Systematics, Taxonomy, and Phylogeny: Glossary
A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z
16S ribosomal RNA (or 16S rRNA) a component of the 30S subunit of prokaryotic ribosomes. It is 1,542 nucleotides in length. Multiple sequences of 16S rRNA can exist within a single bacterium. The 16SrRNA gene is used for phylogenetic studies as it is highly conserved between different species of bacteria and archaea. Carl Woese pioneered this use of 16S rRNA. In addition to these, mitochondrial and chloroplastic rRNA are also amplified.
Adams consensus in cladistic analysis, a type of consensus method that uses the idea that a tree should be thought of as a "set of leaf subset nestings" rather than as a "set of clusters." A group nests within a larger group if the most recent common ancestor of the smaller group is a descendant of the most recent common ancestor of the larger group (from the PAUPDISPLAY Manual). This preserves all nested clades common to a set of source trees (Bininda-Emonds, 2004 - glossary) Adams consensus trees are designed to find the maximum number of components for a given set of cladograms by placing conflicting taxa at the most resolved node common to all the trees. (Forey et al 1992 pp.79-80). Only can be used for rooted trees. Usually preserves more structure than the strict methods, but may show clades in the consensus tree that do not occur in any of the trees in the set, which makes interpretation rather difficult.
AlgorithmIn mathematics and computer science, an effective method (a procedure that reduces the solution of some class of problems to a series of rote steps that give a specific and correct answer) expressed as a finite list of well-defined instructions for calculating a function. Algorithms are used for calculation, data processing, and automated reasoning. (Wikipedia). Modern cladistics and molecular sequencing use algorithms to create cladograms and phylograms
Alpha taxonomythe science of finding, describing and naming species of living or fossil organisms. The term "alpha" refers to alpha taxonomy being the first and most basic step in taxonomy. This science is supported by institutions holding collections of these organisms, with relevant data, carefully curated: such institutes include natural history museums, herbaria and botanical gardens.
Ancestorin this context, an organism, or more correctly a population, lineage, or species, that through evolution gives rise to one or more descendants that generally belong to a distinct taxon or species to itself. The identification of ancestors and descendants is a central aspect of evolutionary systematics. In contrast, cladistics denies it is ever possible to know an ancestor (unless one can actually observe evolution in a laboratory). "No matter how well we understand our group, its taxonomy, paleontology and anatomy, we can never know if one taxon is ancestral to another" (Paraphyly Watch blog - Transitional Fossils, Microbes & Patrocladistics). See also Ancestral group, common ancestor.
ApomorphyIn cladistics, an apomorphy is a unique derived or specialized character trait found in a particular taxon, which is also possessed by a common ancestor. character.
A trait which characterizes an ancestral species and its descendants. This is an evolutionary novelty for the group. These are evidence for the existence of a group. Put another way, attributes shared in common are taken to indicate a shared evolutionary history.
A novel evolutionary trait that is unique to a particular species and all its descendants and which can be used as a defining character for a species or group in phylogenetic terms. Hence, the possession of feathers is unique to birds and defines all members of the class Aves. An apomorphy that is restricted to a single species is termed an autapomorphy. It alone cannot provide any information about the phylogenetic relations of that species, although it can indicate the degree of divergence of a species from its nearest relatives. An example is speech, which is found solely in humans (Homo sapiens) and not in other primates. An apomorphy that is shared by two or more species or groups is termed a synapomorphy. Such traits define the strictly monophyletic groups, or clades, which are the basis of cladistic classification systems (see cladistics). Compare plesiomorphy.
A Dictionary of Biology, Oxford University Press, © Market House Books Ltd 2000
Apomorphy-based taxon (or clade)a group comprising all species descended from a common ancestor characterised by specific apomorphies. Apomorphy-based taxa are rarely used in cladistics because of the difficulty of determining when a particular trait appeared and whether its presence can be reliably determined. In contrast, all Linnaean and Evolutionary systematic taxa are apomorphy-based (either paraphyletic or monophyletic). Contrast with node-based and stem-based
Artin this context we don't mean the Renaissance masters, or the French impressionists, but the role of subjective assessment and intuition in science, a heresy for the advocates of neo-pragmatism and extreme empiricism, but unavoidable if we are to understand something as subtle and complex as the history and nature of life on Earth. I very much like Mike Taylor's comments here on the "How to choose between specific and generic separation" in a Sauropod Vertebra Picture of the Week posting.
At this point, I am reminded of when I used to be on a mailing list for wannabe writers...the best advice I saw on that list was from Jane MacDonald: My personal advice is don't overdo, or underdo, anything in your writing. Do it exactly right. That's my attitude to drawing genus boundaries. It is, frankly, an art; and there are no substitutes for taste, experience, judgement, familiarity with the group in question and all those other touchy-feely qualities that uber-cladists would love to find a way to abolish if they could. But they can't. There is no algorithm for this. I also think of an observation by computer scientist Bjarne Stroustrup, the inventor of the C++ programming language: "Design and programming are human activities; forget that and all is lost." The same is true of palaeontology. (And of, well, everything.)
ASCII Phylogenetic TreeAs here defined, an ASCII phylogeny, or more correctly an ASCII Phylogenetic Tree, is a dendrogramor tree diagram which uses ASCII-text format to draw supertrees. ASCII Phylogenetic Trees might also informally be referred to as ASCII Cladograms, but that is inaccurate because cladograms are, pragmatically speaking, not actually phylogenies but branching diagrams depicting patterns of shared similarities (O'Keefe & Sander 1999) from which evolutionary hypotheses can be constructed. The ASCII Phylogenetic Tree format was created by T. Mike Keesey, who used them to show dinosaur phylogenetic relationships in the old Dinosauricon. Mikko Haaramo adopted this format, but refined it with the introduction of the grave ( ` ) for the corners, for his own phylogenetic archive. This useful format was then adopted on the Dinosaur Mailing List and by paleo enthusiast webmasters like Jack Conrad (The Vertebrate Phylogeny Pages), Justin S. Tweet (Thescelosaurus!), Øyvind M. Padron (The Dinosauria), and Toby White and myself (Kheper palaeo and Vertebrate Notes, and finally Palaeos.com, here). The format has now become pretty standard in any paleo geek text-based phylogenetic diagram.
Available nameA name that is correctly proposed according to the International Code of Zoological Nomenclature. An available name is not necessarily the valid name.
BasalPreferred cladistic substitute for "primitive", as it is felt the latter may carry false connotations of inferiority or a lack of complexity. In cladograms, basal taxa are those terminal taxa that first diverged from the root. The term basal is only be correctly applied to clades (species or higher groups) of organisms, not to individual traits possessed by the organisms. There is however a tendency for terms like basal and stem to appear as rather vague alternatives to "primitive" or "ancestral" in cladistic paleontological literature and especially popularised accounts and comments thereof
Bayesian inferencea method of statistical inference, used for example in phylogenetics, in which some kinds of evidence or observations are used to calculate the probability that a hypothesis may be true, or else to update its previously calculated probability. A form of likelihood analysis that differs from maximum likelihood in that it considers all possible trees (phylograms or cladograms), not just the most likely one, but gives proportionally more credence to the more likely ones. Confidence is measured in terms of probabilities.
Binomial nomenclature Linnaean universal standard of biological scientific notation, according to which every species is given a distinct two-part name. The first part, think of it as like the surname, is the genus, which is capitalised, the second part the species, written completely in lower case, is like the given name. Both names are by convention written in italics (or if that is not possible, underlined, or if even that is not possible say with ASCII text, then there is an underscore character before and after the name, _like this_). So in the case of Tyrannosaurus rex, Tyrannosaurus is the genus (capital "T"), and rex (small "r") the species. Finally, the name of the discoverer of the species is added (if the species has since been given a new genus the discoverer's name is placed in brackets) along with the year of publication of the scientific paper describing that particular species. This usage is not mandatory in popular and semi-technical books, but is when describing or listing species in a technical journal or a Museum. The species name can also be abbreviated by only using the first letter of the genus and a period, after which comes the species name. The species name on its own can be written as T. rex (but never "T-rex", it is not a car!). Any student of natural history will be familiar with this approach. I have noticed however a tendency among paleontologists to give every new discovery a new genus as well as a new species, leading to an over-excess of monotypic genera (each genus only having one species). This was and is exacerbated by the cladistic revolution, where even species previously placed in a genus are moved to their own genus, especially if precise phylogenetic relationship is uncertain (which it almost always is in these cases) only adding to the multiplication of names (Paleo artist and author Greg Paul at one time (Predatory Dinosaurs of the World, 1988) went in the opposite direction, lumping species from even fairly distant genera together; e.g. most dromaeosaurids became Velociraptor, although his more recent work The Princeton Field Guide to Dinosaurs (2010) tends to swing back to less species per genus). There is also a move among several proponents of phylogenetic nomenclature and the Phylocode to abandon the binomial altogether, and emphasise only the phylogenetic relationships (which wouldn't necessarily be evident from the name alone). This would actually only be a small step when considering vertebrate paleontology alone in view of the above mentioned tendency (but not for example Pleistocene mammals which have a very good fossil record!), but would be a nightmare if cataloging or referencing all of the other millions of named and described species. However it is probably unlikely that the Phylocode will become a majority position any time soon.
Bootstrap a sampling method used in phylogenetics. Boostrapping measures how consistently the data support a given tree topology. It does not determine how accurate a cladogram is; it only gives information about its stability, and helps assess whether the sequence data is adequate to validate the topology (branching order).
Bubble diagramInformal neologism used by yours truly for a spindle diagram with rounded rather than angular contours. Also called a romerogram.
CategoryAny rank within the classification hierarchy, e.g., family, subfamily, subspecies.
Change of rankWhen a name is moved from one level of the classification system to another, e.g., when De Lotto (1955) moved Ceroplastes destructor brevicauda from the subspecies to the species rank C. brevicauda this was a change of rank.
Character any recognizable trait, attribute, quality, feature, or property of an organism, of an organism used for recognizing, differentiating or classifying a taxon. used to reconstruct phylogenies. Characters may be morphological, behavioral, physiological, or molecular. In cladistics, the character is thought to be derived and vary from a corresponding feature in a common ancestor of the organisms being studied.
Character Statethe mutually exclusive conditions of a character. (one of the possible alternative conditions of the character). For example, "present" and "absent" are two states of the character "hair" in mammals. Together, characters and character states compose what are termed character statements.
ChronospeciesOne or more species which continually changes from an ancestral form along an evolutionary scale. This sequence of alterations eventually produces a population which is physically, morphologically, and/or genetically distinct from the original ancestors. Throughout this change, there is only one species in the lineage at any point in time, as opposed to cases where divergent evolution produces contemporary species with a common ancestor. Relies on an extensive fossil record, since morphological changes accumulate over time and two very different organisms could be connected by a series of intermediaries. The related term paleospecies indicates an extinct species only identified with fossil material. To avoid unnecessary multiplication of terminology (and paleontology-neontological distinctions) these terms are here synonymised. For example, changes in the Permian lepospondyl amphibian Diplocaulus over time may imply a chronospecies (= paleospecies).
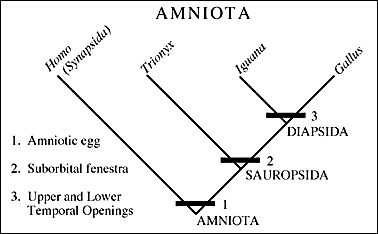 |
| Cladogram showing four species (human, turtle, lizard and bird) and three clades, each defined buy its own synapomorphies (shared unique characteristics). Most cladogram matrixes involve many hundreds of such characteristics
|
CladisticsRigorous methodology first developed by Wili Hennig, to evaluate and reconstruct phylogenetic hypotheses. The results of cladistic analyses are often represented in the form of a branching diagram, called a cladogram. It is important to keep in mind that cladistics is not the same as phylogeny, and cladograms are not phylogenetic diagrams of ancestor-descendant relationships! As with Linnaean classification, cladistics provides a nested hierarchy where an organism is assigned a series of names that more and more specifically locate and define it within the hierarchy. However, unlike Linnaean classification, phylogenetic classification only allows monophyletic clades, excludes both paraphyletic and polyphyletic groups. It also does not assign ranks (e.g. class, phylum) to the hierarchical levels.
The late 1970s and early 1980s saw conflict between the two schools of Pattern Cladism and Hennigian systematics, although this has since been resolved, and cladistics today is generally based equally on both. In the 1980s and 90s cladistics became the dominant paradigm in biological systematics, supplanting the previous Linnaean-based evolutionary taxonomy in all fields except botany. Together with molecular phylogeny it forms the current Phylogenetic paradigm.
With the advent of powerful computation, cladistics has come to use statistical procedures such as Bayesian analysis and Maximum Likelihood. Such computer-based cladistics are now used especially in paleontology to determine the relationships between various fossil organisms. Often relying on supermatrixes and incorporating large numbers of species and hundreds of character states, they tend to gives very different results to that of earlier hand-coded diagrams, which emphasied instead a smaller number of well-recognised synapomorphies.
Despite being often referred to as "phylogeny", cladistics today does not seek to describe the actual course of phylogeny in deep time a la evolutionary systematics, but only to select the most viable hypothesis of phylogenetic relationships, given the available data (empirical method).
It is held by cladists that taxa (if recognized) must always correspond to clades, united by apomorphies (derived traits) which are discovered by a cladistic analysis. To this end, cladistics collects character data only from the taxa being studied, and do not consider the inferred characters of ancestors.
CladogenesisThe division of an ancestral parental lineage into two or more daughter lineages or species. At one time, cladogenesis was recognised, along with anagenesis, as one of the two types of gradual evolution. Because the highly formalised trees that cladistics use on do not show anagenesis, a misplaced literalism led to cladogenesis, in the sense of the division of a common ancestor into two daughter species, being accepted as the standard form of speciation. However, other evolutionary processes, especially budding and merging, involve asymmetrical divergence and therefore paraphyly.
CladogramA dichotomous phylogenetic tree that branches repeatedly, suggesting a classification of organisms based on the sequence in which evolutionary branches arise; a nested diagram of synapomorphies indicating possible relations between groups; each point of branching represents divergence from a hypothetical common ancestor. Cladograms only give information about branching order, not about the amount of change or difference (unlike phylograms), the diversity of each taxon (unlike spindle diagrams) or stratigraphic range (unlike chronogram, although a cladogram can also be drawn as a chronogram).
In the 1980s, cladograms used Hennigian methodology and were based on immediately apparent synapomorphies. From the 1990s onwards, computational phylogenetics began to be used in the generation of cladograms, and these have now long since become standard. Characters pertaining to each taxon are run through computer algorithms to determine phlogenetic relationships. Although traditionally cladograms were generated largely on the basis of morphological characters alone, nowadays DNA and RNA sequencing data have been used as well. All have different intrinsic sources of error. For example, character convergence (homoplasy) is much more common in morphological data than in molecular sequence data, but character reversions that are unrecognizable as such are more common in the latter (see long branch attraction). Morphological homoplasies can usually be recognized as such if character states are defined with enough attention to detail. The researcher must decide which character states were present before the last common ancestor of the species group (plesiomorphies) and which were present in the last common ancestor (synapomorphies) and does so by comparison to one or more outgroups. The choice of an outgroup is a crucial step in cladistic analysis because different outgroups can produce trees with profoundly different topologies. Note that only synapomorphies are of use in characterizing clades.
Classification. In biology, a classification is a system of uniting taxa into a system of interconnected units in order to reflect features uniting them. Classifications may be either artificial (built on arbitrarily-chosen features to facilitate the worker's convenience) or natural (supposedly derived from the evolutionary relationships of the taxa). Most authors would currently favour the latter, though artificial classifications may still be in use for groups of organisms (such as anamorphic fungi) in which evolutionary relationships are difficult to establish. Many groups of organisms may have different classificatory systems in use at the same time due to differing opinions between different authors, and classifications may also change as authors refine their investigations. Classification should be distinguished from nomenclature, which is the investigation of correct names for taxa. Classification and nomenclature together form taxonomy.
Coalescent TheoryA method for comparison of gene sequences in populations to find the most likely common ancestor sequence.
Cohesion Species Concept defines species as the most inclusive group of organisms having the potential for genetic and/or demographic exchangability
Common ancestorThe ancestral species that gave rise to two or more descendant lineages, and thus represents the ancestor they have in common, and from which later species and groups evolved. The idea of a common ancestor is central to evolutionary thinking from Darwin onwards. In the Modern Synthesis' Evolutionary Systematics the common ancestor is usually shown as the most suitable fossil form at the base of a lineage, where it may or (more likely given the small number of species known from those which actually lived in past ages) or may not be an actual ancestor, more often it is a sort of grand-uncle rather than grandfather). Evolutionary Systematics is based on identifying and determining the actual traits of an ancestral species or, more usually, supra-specific taxa.
In an attempt to establish greater rigour and precision, Cladistic phylogeny defines the most recent common ancestor as the originator of a clade; in other words the first species or organism to possess the unique attributes of that clade. Contrary to popular opinion, cladograms do not actually show the actual common ancestor; such an organism or group would be by definition paraphyletic, and hemnce automatically forbidden by cladistic logic. Cladistics therefore rejects the possibity of knowing the actual common ancestor, and instead posits a hypothetical common ancestor. However, a basal taxon may have some features in common witrh the common ancestor.
 |
| A Consensus Cladogram
|
Consistency index (CI)In cladistics, the measure of the parsimony fit of a character to a tree, or of the average fit of all characters to a tree. Varies from 1.0 (perfect fit) to a value asymptotically approaching zero (poorest fit). It is inflated by autapomorphies which can only take the value 1.0; thus a totally uninformative data set (consisting only of autapomorphies) could return a CI equal to 1.0. Compare retention index. (Michael D. Crisp - Introductory glossary of cladistic terms). The per-character consistency index (ci) is defined as m/s, where m is the minimum possible number of character changes (steps) on any tree, and s is the actual number of steps on the current tree. This index hence varies from one (no homoplasy) and down towards zero (a lot of homoplasy). The ensemble consistency index CI is a similar index summed over all characters.
Cotype (Zoological Code) this term has been used in the past to refer to either a paratype or a syntype. Its use is now discouraged.
Crown groupin cladistics, a group consisting of living representatives, their ancestors back to the most recent common ancestor of that group, and all of that ancestor's descendants. The name was given by Willi Hennig as a way of classifying living organisms relative to extinct ones. Though formulated in the 1970s, it was not commonly used until its reintroduction in the 2000s. The usual definition of a crown group is the smallest monophyletic group, or "clade", to contain the last common ancestor of all extant members, and all of that ancestor's descendants. Extinct side branches on the family tree will still be part of a crown group. For example, if we consider the crown-birds (i.e all extant birds and the rest of the family tree down to their last common ancestor), extinct side branches like the dodo or great auk are still descended from the last common ancestor of all living birds, so falls within the bird crown group.
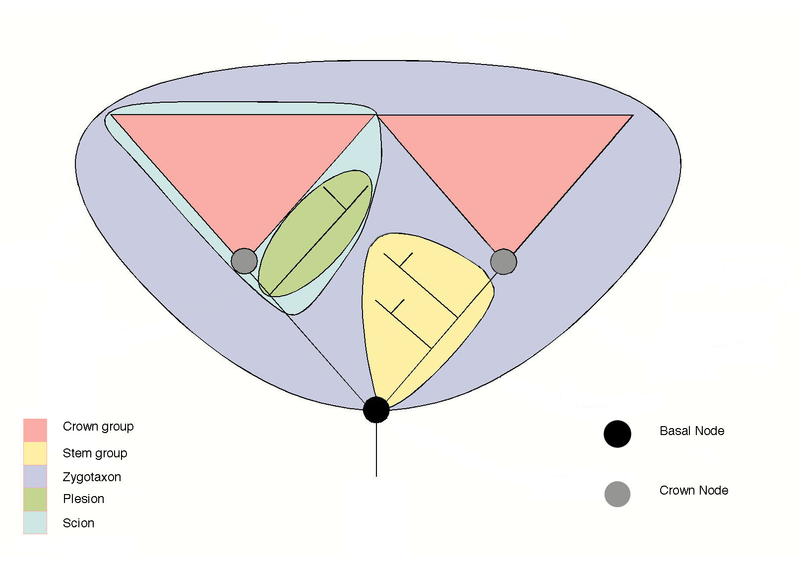 |
| The stem and crown group concept. The two pink groups represent a pair of crown groups, the last common node of which is the basal node. Terminology is from Craske, A. J. and Jefferies, R. P. S. (1989) A new mitrate from the late Ordovician of Norway, and a new approach to subdividing a plesion. Palaeontology 32, 69–99 and Budd, G. E. (2001) Tardigrades as "stem-group" arthropods: the evidence from the Cambrian fauna. Zoologischer Anzeiger 240, 265-279. The diagram shown here is revised from the original to clarify that the stem group does not include the basal node (ancestor) of the crown group. For explanation of terminology see Wikipedia - Crown Group page.
From Wikipedia. Diagram and text by Graham Budd. Revised by Peter Coxhead. |
Decay indexIn cladistic analysis, the number of additional steps required to dissolve a given clade. (see also Bremer support)
Michael Allaby, 1999, Dictionary of Zoology
DendrogramThere doesn't seem to be an agreed meaning of this term. Michael Crisp's cladistic glossary defines it as any branching diagram or tree, such as a cladogram. Mayr & Bock 2002(for the evolutionary systematics camp) contrast the "Hennigian cladogram" with the "Darwinian dendrogram". The Wikipedia page gives another definition again: "a tree diagram frequently used to illustrate the arrangement of the clusters produced by hierarchical clustering. Dendrograms are often used in computational biology to illustrate the clustering of genes or samples." As used on Palaeos, a dendrogram is any informal phylogenetic cladogram-like diagram, a sort of composite of published trees, or simply the author in question's random opinion.
Derivedsame as apomorphy; a derived character / trait is inferred to be a modified version of a more primitive condition of that character and therefore inferred to have arisen later in the evolution of the clade.
Diagnosis: statement in words that purports to give those characters which differentiate a taxon from other taxa with which it is likely to be confused.
ElectrophoresisThe method of distinguishing entities according to their motility in an electric field. In evolutionary biology and molecular sequencing, it has been mainly used to distinguish different forms of proteins. The electrophoretic motility of a molecule is influenced by its size and electric charge.
EmendationIn taxonomy, an intentional change to a previously proposed name, e.g., Lindinger proposed the emendation Hemiberlesea for the armored scale Hemiberlesia indicating that it was originally improperly formed.
Epitype (Botanical Code) a specimen designated at a later date to characterise a species, where the original type material is not sufficient to do so. The original type retains name-bearing status, and should the epitype later prove not to be conspecific, the name remains with the holotype (however, it is not uncommon for the International Association of Plant Taxonomy to conserve the common understanding of a name by setting aside the holotype in favour of the epitype).
 |
| Elvis Presley in concert, from the 1973 television broadcast Elvis: Aloha from Hawaii.
|
Elvis taxona taxon which has been misidentified as having re-emerged in the fossil record after a period of presumed extinction, but is not actually a descendant of the original taxon, instead having developed a similar morphology through convergent evolution. This implies the extinction of the original taxon is real, and the two taxa are polyphyletic. The term was coined by D. H. Erwin and M. L. Droser in a 1993 paper to distinguish descendant from non-descendant taxa: "Rather than continue the biblical tradition favored by Jablonski [for Lazarus taxa], we prefer a more topical approach and suggest that such taxa should be known as Elvis taxa, in recognition of the many Elvis impersonators who have appeared since the death of The King." Lobothyris subgregaria, a brachiopod from the early Jurassic period, is one example of such a taxon. By contrast, a Lazarus taxon is one which actually is a descendant of the original taxon, and highlights missing fossil records, which may be filled later. A Zombie taxon is a taxon sample that was mobile in the time between its original death and its subsequent discovery in a site of younger classification, like, for example, a trilobite that gets eroded out of its Cambrian-aged limestone matrix, and reworked into Miocene-aged siltstone.
 |
|
|
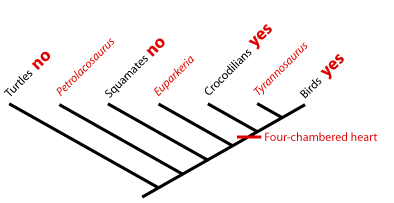
Extant Phylogenetic Bracket, Phylogenetic bracketingIn 1999, Larry Witmer described how unknown character states for fossil taxa are reconstructed with respect to extant taxa called the extant phylogenetic bracket (EPB). This is the bracket formed on either side of the taxon with the missing information by extant taxa in which the character state is known. Using it, we can make three types of inference, listed in order of decreasing confidence. Consider the distribution of a soft-tissue character - the four-chambered heart - among three fossil reptiles:
-
Type I Inference: Tyrannosaurus is bracketed by birds and crocodilians, both of which have the derived character. With no contrary positive evidence, the simplest assumption is that Tyrannosaurus had it also.
-
Type II Inference: The basal archosauriform Euparkeria is bracketed by crocodilians and squamates. Crocs have the derived character, squamates don't. Thus, we are much less secure than above in inferring it in Euparkeria , but presence of some sort of hard tissue correlate of that trait might increase our confidence.
-
Type III Inference: The basal diapsid Petrolacosaurus is bracketed by squamates and turtles, neither of which have the derived character. Our confidence in its presence in the extinct form is very low. We would need strong positive fossil evidence to argue for its presence.
"Gap codings" this is not a formal term but refers to the situation in cladistics, when a 'daughter' character is logically dependent upon the state of a 'parent', and cannot be coded when the parent is absent. For example, the position of the frontal appendage in an arthropod can only be coded in taxa that possess a frontal appendage in the first place. In morphological analyses, this assigns double weight a priori to absences in the 'parent' character (because the daughter is always contingent, that is, dependent on the parent character), and can artificially inflate support for particular clades, and hence affect overall tree topology. This situation is hard to avoid when selecting characters across a range of fossils, which include taxa with unusual or differing morphologies. In analyses of nucleotide data the situation is different, because gaps may be the result of shared deletions from an ancestral sequence and hence be informative.
"Garbage in, garbage out"self-explanatory phrase borrowed from computer programming. If the characters used in phylogenomics (and cladistic analysis in general) are unreliable, even the most accurate tree reconstruction method can fail. Therefore, methods focusing on the most reliable characters have been developed in order to reduce the impact of inconsistency.
GenealogyTerm derived from Greek γενεά, genea, "generation"; and λόγος, logos, "explanation". The study of families and the tracing of their lineages and history. Genealogists use oral traditions, historical records, genetic analysis, and other records to obtain information about a family and to demonstrate kinship and pedigrees of its members. The results are often displayed in charts or written as narratives. In evolutionary thought, such as cladistics, due to the alternate translation of γενεά as "race" the term can be used as a synonym for phylogeny.
Genetic AlgorithmsComputational systems based upon an implementation of natural selection as an algorithm for classification or optimization.
GenusThe taxonomic rank between family or tribe and species, and used to define group of closely related organisms that differ in only very minor ways. In the Linnaean system of binomial nomenclature, the genus is written in italics, with a capital letter, in front of the species name, or on its own. e.g. with Tyrannosaurus rex, the name Tyrannosaurus is the genus, and T. rex (no hyphen!) is the species. Used in evolutionary systematics; in cladistic classification every genus is only allowed two species (because of excessive formalism regarding cladogenesis), and Linnaean genera are always oversplit and new names created, resulting in much taxonomic confusion (for example in paleontology the established dinosaur genus Iguanodon has been split into about a dozen different monospecific genera (link). See also the discussion at Sauropod Vertebra Picture of the Week. It may be that the Phylocode will discard binomial nomenclature altogether (although there is obvious resistance to this).
Ghost lineagein cladistics, a phylogenetic lineage that is inferred to exist, for example by matching a cladogram against geological time, but is not known from the fossil record.
When we know that two taxa are sister taxa (descendants of the same recent common ancestor), we in essence know that they originated at the same point in geologic time - the time of their last common ancestor and the speciation event that gave rise to them. Say we know one of these taxa from 100 million year old rocks, and the other from 90 million year old rocks. Even without seeing a fossil, we know that the second group must have representatives dating back at least to 100 million years, simply from its sister-taxon relationship with the other. A lineage like this, whose existence can be inferred from the cladogram, but which is not known from actual fossils is called a ghost lineage. The examination of ghost lineages should allow biostratigraphers to refine their models of the stratigraphic ages of organisms.
Gradism; Gradisticsas used here, the opposite (or complement) of cladistics; understanding phyloigeny in terms of evolutionarey tramnsformation and ancestor-descendent relationships. Includes the evolutionary systematics of .
Great chain of beingmetaphysical premise, popular from the classical world until the early 19th century, that all beings constitute a single continuous series of forms in an unbroken gradation from God through countless intermediate spiritual and material stages to formless matter. Also used more profanely to justify feudalism, the Church, etc. With the development of naturalistic theories of evolution, representations of the great chain of being were replaced by phylogenetic trees and secular cosmology. Nevertheless the meme of great chain of being remains popular outside science, including philodsophical and pop cultural referneces to "ascent". See also The March of Progress.
Hapantotype (Zoological Code) for protists with complex life cycles (such as Apicomplexa), a series of specimens taken from different stages of the life cycle acting as the type. Though composed of multiple specimens, a hapantotype series is treated as a single holotype, and a lectotype may not be designated from within it. Should a hapantotype turn out to contain specimens from more than one species, specimens may be excluded from it until only conspecific ones remain.
Heterotachy. Variation in the evolutionary rate of a given position of a gene or protein through time. Can lead to phylogenetic reconstruction artefacts where unrelated taxa have converged in their proportions of invariable sites. Unlike other types of bias, heterotachy does not leave any evident traces in sequences, and therefore are particularly difficult to detect.
Holotypein taxonomy, a single specimen (or illustration for the Botanical Code) designated by the author in the original publication. Under the Zoological Code since 1999, any species description that does not explicitly designate a type is deemed invalid, and the species name a nomen nudum.
Holophyletic, HolophylyAshlock 1971 coined the term to resolve the ambiguity between the Haeckelian (evolutionary systematic) and Hennigian (phylogenetic systematic, cladistics) definitions of monophyly, and that usage is followed here. Refers specifically to the definition that a group contains the common ancestor, all organisms descended from the common ancestor, and no other organisms. The term has not gained widespread acceptance in the scientific community, probably because monophyletic is so widely used and has the same meaning.
HomonymOne of two or more scientific names that are identical but pertain to different organisms, e.g., Eriococcus mancus Ferris, 1955 and Eriococcus mancus (Maskell, 1897); Onceropyga Ferris, 1955 and Onceropyga Turner, 1904.
International Code of Zoological Nomenclaturewidely accepted convention in zoology that rules the formal scientific naming of organisms treated as animals. The rules principally regulate:
-
how names are correctly established in the frame of binomial nomenclature,
-
which name has to be used in case of conflicts among various names,
-
how names are to be cited in the scientific literature.
The rules and recommendations have one fundamental aim: to provide the maximum universality and continuity in the scientific naming of animals. The code is published by the International Commission on Zoological Nomenclature (ICZN), an organization dedicated to "achieving stability and sense in the scientific naming of animals". The rules in the Code determine what names are valid for any taxon in the family group, genus group, and species group. It has additional (but more limited) provisions on names in higher ranks. Several cladists have argued that the Linnaean based ICZN code needs to be replaced by a new cladistically-based system, the Phylocode.
Infrasubspecificcategory or name of lower rank than subspecies, and, therefore not subject to regulation by nomenclatural Codes; e.g. form, race, variety.
Intuitionin this context, arriving at a scientific (or any creative) hypothesis through a leap of insight. For example, Einstein discovered Special Relativity by imagining what it would be like to ride on a photon. From another perspective, gut-feelings, hunches, creativity, and more. See also art. In systematics, advocates of Phenetics and Cladistics argue on pragmatic grounds that evolutionary systematics should be rejected because it is too "intuitive", and not sufficiently verifiable. However their use of quantitative empirical data without intuition meant they were not able to distinguish homology from homoplasy. Hence all science will always include some intuition and subjectivity.
Isotype (Botanical Code) a specimen deriving from the same individual as the holotype (for instance, a second cutting from the same tree).
Jackknife Value(to be added)
Junior synonymgiving a new name to a species, supra-specific taxon, or clade which already has a scientific name. As a standard, the first applied name is the one that is used in biological and paleontological systematics. Junior synonyms are redundant and hence usually rejected in scientific nomenclature; the exception being when the more recent name is so well known that to change it would cause confusion. For example, the first named fossil which can be attributed to Tyrannosaurus rex consists of two partial vertebrae found by Edward Drinker Cope in 1892 and named Manospondylus gigas. It was only later realised that they belong to the same animal. In this case, the newer name, Tyrannosaurus rex (named by Henry Fairfield Osborn in 1905) was retained, and the older one Manospondylus gigas, rejected. If there are only two synonyms, the most recently described one is the junior synonym; if there are more than two synonyms, the junior synonyms are all but the oldest described one which is the senior synonym.
Junior homonymIf there are only two homonyms, the junior homonym is the most recently described homonym; if there are more than two homonyms, the junior homonyms are all but the oldest described homonym which is the senior homonym, e.g., Eriococcus mancus Ferris, 1955 is the junior homonym and Eriococcus mancus (Maskell, 1897) is the senior homonym.
Justified emendationAn emendation that is correct according to the International Code of Zoological Nomenclature, e.g., the name susani is proposed as a patronym for a woman named Susan; according to the Code the name must be changed to susanaeand is a justified emendation.
Lapsus calumni(abbrev. l.c.) slip of the pen, an accidental mispelling; especially common with some of those difficult latin names. e.g. Poecilopleuron, Poecilopleurum, Poicilopleuron, and Poikilopleuron are all mispellings of the Jurassic theropod Poekilopleuron.
Last (or Latest) Common Ancestor (LCA) the most recent common ancestor of any two (or more) species, which is another way of saying it is the earliest member of a particular clade that includes those species but not more distantly related species. So there are still earlier ancestors, and they would also be common ancestors, but they would include other taxa as well as those being studied, and would stand at the base of a more inclusive clade. e.g. the most recent common ancestor of a dog, a cow, a human and a chimpanzee (Boreoeutheria) is also the common ancestor of a human and a chimp (Hominidae), but it isn't the most recent one (it lived much earlier and evolved into far more groups of animals). Therefore, to limit study to the group or clade under consideration, only those members in that clade, and their most recent (not oldest) common ancestor is considered.

Lazarus taxona taxon that disappears from one or more periods of the fossil record, only to appear again later. An example is Lazarussuchus, an Oligocene member of a clade of freshwater reptiles (Choristodera) thought to have gone extinct at the end of the Mesozoic. As Lazarussuchus is thought to be outside the clade including other choristoderans, it may indicate a ghost lineage going back to the Late Triassic, a span of over 170 million years. There are also examples of "Burgess Shale type fauna", best known from the Early and Middle Cambrian periods, but which, since 2006, have been found in rocks from the Ordovician, Silurian and Early Devonian periods, in other words up to 100 million years after the Burgess Shale (Kühl et al 2009; Siveter et al 07). The term "Lazarus taxon" refers to the account in the Gospel of John, in which Jesus raised Lazarus from the dead. Lazarus taxa are observational artefacts that appear to occur either because of (local) extinction, later resupplied, or as a sampling artefact. If the extinction is conclusively found to be total (global or worldwide) and the supplanting species is not a look-alike (an Elvis species), the observational artefact is overcome. The fossil record is inherently imperfect (only a very small fraction of organisms become fossilized) and contains gaps not necessarily caused by extinction, particularly when the number of individuals in a taxon becomes very low. If these gaps are filled by new fossil discoveries, a taxon will no longer be classified as a Lazarus taxon. A subtle difference is sometimes made between a "living fossil" and a "Lazarus taxon". A Lazarus taxon is a taxon (either one species or a group of species) that suddenly reappears, either in the fossil record or in nature, while a living fossil is a species that (seemingly) hasn't changed during its very long lifetime. Sometimes however, the two are confused or conflated, as with the coelacanth, which is also called a "living fossil" because it was thought to be extinct for tens of millions of years, but then discovered alive.
Lectotypea specimen selected from a syntype series to become the single name-bearing type of the species in order to confirm the identity of the species. The other previous syntypes become paralectotypes.
LengthThe length, or number of steps, is the total number of character state changes necessary to explain the relationship of the taxa in a tree. According to the principle of parsimony, the fewer number of character state changes required, the more likely the tree. A tree with a lower length has less homoplasies and so fits the data better than a tree with a higher length. The tree with the lowest length assumes fewer homoplasies and hence is more parsimonious, and so represents the hypothesis of taxa relationship that is selected.
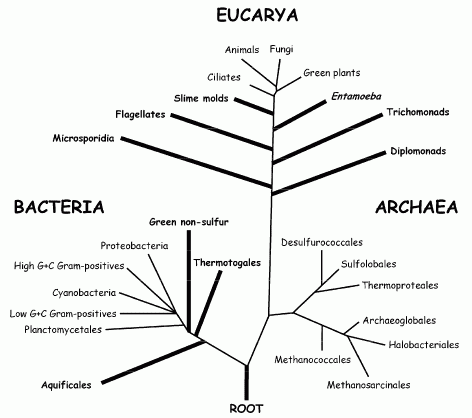 |
The classical view of the universal tree of life, topology inspired from Stetter 1996, mainly based on rRNA comparison. Branches that could be affected by long branch attraction artefacts (e.g., the placement of the root in the bacterial branch or the early emergence of hyperthermophilic taxa amongst bacteria) are given as thick lines.
|
Linnaean classificationhierarchical taxonomy developed by the 18th century Swedish botanist Carl von Linné, (Linnaeus). It was the first systematic classification of life on Earth, in which every species is given it's own binomial designation. So for example anatomically modern human beings are Homo sapiens, genus (the "family name") Homo and species (the specific name) sapiens. In contrast, Neanderthal man is Homo neanderthalensis. Linnaean classification provides a nested hierarchy of levels, each with its own specific characteristics. In this way any organism or species is grouped more and more specifically within the hierarchy. The Linnaean system was originally static, being based on creationism. In the 19th century, applied to the evolution of life and the modern synthesis it became evolutionary systematics, and was used to construct phylogenetic trees. Still foundational to modern biology, Linnaean classification is in the process of being superseded by phylogenetic hypothesis-based cladistic systematics. This latter, with its indefinite series of nested clades, lacks the categorical simplicity and ease of use of the old Linnaean system. Some attempts have been made to integrate the two, but the incompatible methodologies mean that so far these have not been very successful.
Long branch attraction (LBA)A phenomenon in molecular phylogenetic analyses, especially those employing maximum parsimony. Unrelated species or lineages sharing rapid evolutionary rates are artefactually grouped together and hence considered closely related, regardless of their true evolutionary relationships. In other words, unrelated lineages may group on the basis of convergent changes rather than homologies, the long branches being attracted to each other because of chance similarities. For example, in DNA sequence-based analyses, the problem arises when sequences from two (or more) lineages evolve rapidly. For example, rRNA evolutionary rates may vary by a factor of 100 among planktonic foraminifers. As there are only four possible nucleotides, when DNA substitution rates are high, the probability that two lineages will evolve the same nucleotide at the same site increases. When this happens, parsimony erroneously interprets this homoplasy as a synapomorphy (i.e., evolving once in the common ancestor of the two lineages). In phylogenies rooted by a distant outgroup, unrelated fast evolving ingroups will emerge independently as the deepest offshoots, being attracted by the long branch of the outgroup. LBA artefact currently represents a major concern to phylogeneticists, as it is believed to affect the position of virtually every deep-branching lineage. As a result, many organismal relationships in the universal tree, shown as bold lines in the diagram on the right, should be regarded as suspect (note: this particular topology has since been corrected by more recent revisions) This problem can be minimized through improved models of sequence evolution and by using methods that correct for multiple substitutions at the same site, through increased or modified taxonomic sampling and by breaking up long branches adding taxa related to those with the long branches or by using alternative slower evolving traits. Long branche attraction is also a problem with morphology-based cladistics because each branch may have so many unique modifications that tracing shared (ancestral) conditions may be difficult.
Majority rule consensusin cladistic analysis, a consensus method that preserves all relationships appearing in 50% of the source trees. This method allows a group to appear in the consensus even if some of the trees in the set contradict it, as long as a majority of the trees (generally half or more) support the grouping. In fully resolved majority rule consensus, these can appear in the consensus solution so long as they do not contradict relationships that occur more frequently. When comparing only two trees, this method is equivalent to the strict consensus method.
Matrixtabulated data of the characters of all of the taxa used in a cladistic analysis, arranged in rows (taxon) and columns (character). "0" indicates that a character is absent, "1" that it is present. If there are more than one possible character states, these are inicated by further numbers, such as 2 or 3(very rarely more). if the character state is not known (common in the case of fossils, epsceially fragmentary ones), a question mark is used instead. The tabulated data is used to form phylogenetic hypotheses, which can be diagrammatically represented as cladograms.
Molecular clockthe premise that the rate at which mutational changes accumulate is constant over time. The difference between the form of a molecules in two species is then assumed to be proportional to the time since the species diverged from a common ancestor, and molecules can be used to date the tree of life. In the late 1960s, the neutral theory of molecular evolution provided a theoretical basis for the molecular clock, though both the clock and the neutral theory were controversial, since most evolutionary biologists held strongly to panselectionism (Adaptationism), with natural selection as the only important cause of evolutionary change. (Wikipedia, etc). Although subject to certain caveats and continuing debate, the notion of the molecular clock has proven to be an important and useful tool in many contexts Searls, 2003 glossary The tendency now is to calibrate the molecular clock by the fossil record (Donoghue & Benton 2007). Earlier problems associated with this method for example, the evolution of animal phyla during the Precambrian (early in the Proterozoic (ref), for which there is absolutely no fossil evidence) have since been largely rectified. Even so, it is difficult to believe that the molecular clock rate does not vary greatly at particular times, for example accelerating during periods of rapid evolutionary radiation (the Cambrian explosion in this example). In other instances evolution may be more constant, and molecular clocks more reliable. The choice of molecule used may also be significant (reference to be included).
Molecular phylogeny, Molecular systematicsUse of data from informational macromolecules (DNA, RNA, and/or proteins) as characters for phylogenetic analyses in order to map out the evolutionary tree of life. That is, the use of the structure of molecules to gain information on an organism's evolutionary relationships. Includes methods based on overall similarity (Phenetics), like electrophoresis, immuno-distance and DNA-DNA-hybridisation, as well as methods that are based on parsimony, like restriction-site-analysis and sequencing sequencing). Generally speaking, the more closely related two organisms are, the more similar their gene sequences will be. By statistically comparing the similarities and differences in the sequence between the same gene from various organisms, we can deduce the pattern of how those organisms are related, and shown in a phylogram. Despite the similarities (both involve dichotomous branched trees), these are not cladograms. Over the last decade or so, molecular phylogeny has supplanted morphology-based cladistic as the primary way of understanding the evolution of life on Earth, giving rise to phylogenetics, the synthesis of molecular phylogeny and cladistics, based on a total evidence approach and supermatrix trees. .
Monotypicin Linnaean classification, a higher ranked taxon that contains only a single species. e.g. Ginkgo is a monotypic genus that contains a single extant species, biloba; the family Ginkgoaceae is similarily a monotypic family. In cladistics (and especially vertebrate paleontology), allowing only the type species in that genus; all other species are given their own genera. This is in keeping with a phylocode approach (which rejects supra-specific taxa such as genera, families, phyla etc), and understandable especially when dealing with fossil taxa where there is only very limited information (sometimes all that is known of a species are a few scraps of bone) and phylogenetic placement is uncertain.
Morphology
The gross form and structure of an organism, or of a part of an organism. In paleontology and phylogenetic analysis may refer to the form or structure of a particular bone or shell, and its comparison with that of similar species; phenomic traits.
Short for Morphology-based phylogeny (see next entry), and also referring to any resulting phylogenetic trees that may be derived from this methodology. One of the two rival phylogenetic methods currently in use , the other being molecular phylogeny. Although the latter has become the default paradigm (e.g. phylogenetics), here at Palaeos we have also given equal weight to morphology-based approaches. Morphology-based phylogeny is more or less synonymous with traditional cladistics, newer total evidence cladistic approaches also incorporate molecular sequencing data although the results may sometimes be a little strange (such as pleurodire turtles as highly derived crown group cryptodires)
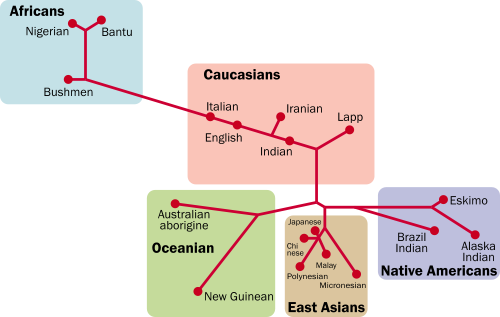 |
| This genetic distance map made in 2002 is an estimate of 18 world human groups by a neighbour-joining method based on 23 kinds of genetic information.
|
Neighbor-joininga bottom-up clustering method for the creation of phenetic trees (phenograms), created by Naruya Saitou and Masatoshi Nei. Usually used for trees based on DNA or protein sequence data, the algorithm requires knowledge of the distance between each pair of taxa (e.g., species or sequences) in the tree. (Wikipedia. It works something like this. Locate the pair of sequences that are the most similar, and treat this as a single avergaed pair. Match it with the next most similar and so on, to build up a tree of successive nested groups. Not as computation-heavy as other methods. However, because neighbour joining trees do not convey phylogenetic information they have been replaced by parsimony and likelihood statistical methods.
Neotypea new type specimen designated subsequent to the original description. A neotype can only be designated if a type was not originally designated (for species published before 1999), or if the original type(s) is lost or destroyed. In a very few cases in zoological nomenclaure (such as for Coelophysis bauri) a neotype has been designated to replace an unidentifiable holotype - such an action, however, requires a Decision by the ICZN.
New combinationWhen a species is transferred to a different genus for the first time.
Non- As phylogenetics does not allow the use of paraphyletic or ancestral taxa, it becomes difficult to refer to groups at the base of any evolutionary lineage. One way is to use prefixes like basal and stem, but these can tend to fuzzy vagueness, e.g. "basal archosauria" is not a correct term for all "thecodonts" but only strictly speaking refers to the most basal node or taxon of clade Archosauria. In this context, stem would be more accurate, but seems to be less often used. Another method is to use non-. For example, because the monophyletic clade Dinosauria includes not just dinosaurs but birds (because, cladistically speaking, birds are dinosaurs) dinosaurs as traditionally defined are not called dinosaurs but non-avian dinosaurs. The ancestors of dinosaurs, such as lagosuchids and silesaurids, then become non-dinosaurian dinosauromorphs. This problem does not arise in evolutionary systematics, which recognises and identifies ancestral groups.
Numerical taxonomysame as phenetics; a method of generating phylogenies that is based on large numbers of quantifiable (measurable) characters which groups organisms with respect to overall similarity.
Nodeany point in a cladogram where branches diverge or end. In cladistics, nodes of phylogenetic trees represent taxonomic units. Internal nodes (or branches) refer to hypothetical ancestors whereas terminal nodes (or leaves).External nodes, which are at the end of a each branch represent terminal taxa, generally extant species but where paleontological data is considered they can also include fossil species. Internal nodes are where a single ancestral lineage breaks into two or more descendant lineages. In rooted trees, internal nodes represent hypothetical common ancestors.
Nomen conservandum (abbreviation nom. cons., plural nomina conservanda – latin for "a name to be preserved") A nomen conservandum is a name that, under strict application of the appropriate code of nomenclature, should be invalid, but which the relevant commision has decided should be upheld in the interests of stability and communication. This may, for instance, involve the preservation of a well-known name for a taxon rather than its otherwise mandatory replacement with an unfamiliar or poorly-defined senior synonym. To what extent a name is conserved depends on the case - a name can be universally conserved, so that it takes priority over any non-conserved synonym, whether already known or recognised later, or it may only be conserved relative to the specific name(s) recognised in competition at the time.
For instance, the name Meganthropus africanus was established for a fossil hominid by Weinert in 1950. Later, this was synonymised with Australopithecus afarensis Johanson et al., 1978 within the genus Australopithecus. As there is already an Australopithecus africanus Dart, 1925, A. afarensis was the correct name. However, some authors have suggested that Australopithecus afarensis should be removed from Australopithecus and placed in the genus Praeanthropus. As the homonymy with Australopithecus africanus would then be removed, the technically correct name for the species would then be Praeanthropus africanus (Weinert, 1950). However, a request was made to the International Commission on Zoological Nomenclature for the preservation of the species name afarensis (nomen conservandum) due to its high public profile, and to prevent confusion with the equally well-known Australopithecus africanus. The ICZN upheld this request in 1999, meaning that even when placed in a different genus, Australopithecus afarensis remains afarensis.
Nomen dubium (abbreviation n. d., plural nomina dubia) A nomen dubium (Latin, "doubtful name") is a taxon that has not been characterised in enough detail and whose type material is not sufficient for it to be identified. For instance, a number of dinosaur taxa named in the 1800s such as Trachodon were based on isolated teeth. Unfortunately, teeth in reptiles do not generally differ between species, meaning that fossilised teeth usually cannot be reliably identified to a particular species.
The significance of a taxon being declared a nomen dubium is often misunderstood. Contrary to popular belief, a nomen dubium is not invalid, in the way a nomen nudum is. A nomen dubium is still available for consideration in terms of synonymy and/or homonymy, and if a name previously regarded as a nomen dubium is able to identified with a better distinguished taxon that was named later, the nomen dubium is still the senior synonym, and hence the correct name for the taxon. One well-known example of this involves Allosaurus fragilis Marsh, 1877, which was suggested in the past as synonymous with Antrodemus valens Leidy, 1870, and Allosaurus appeared as Antrodemus in a number of older sources. However, Antrodemus is based on a single isolated tail bone, which is not sufficient to characterise the species. Allosaurus is currently regarded as a valid taxon, but this is because Antrodemus cannot be conclusively identified with it, not because Antrodemus is a nomen dubium. See New papers in Geobios (and nomenclatoral gripe) and follow-up messages on the Dinosaur Mailing List for an example of an argument on the appropriate application of a nomen dubium.
Nomen nudum (abbreviation: n. n., plural nomina nuda) A nomen nudum (Latin, bare name) is a name that fails to meet the requirements for being validly published under the appropriate code of nomenclature (for instance, no published description). A nomen nudum has no official nomenclatorial standing, and does not compete for synonymy, homonymy, etc. Should a name that was previously a nomen nudum ever be validly published, its priority dates from valid publication, not from original appearance.
In these days of the internet and widespread media, nomina nuda are sometimes a significant issue (especially in vertebrate palaeontology). It is not uncommon for significant and/or interesting discoveries to be popularised in newspapers, newsgroups, etc. before their appearance in the professional literature. Any names that appear in such formats are generally nomina nuda.
Nomen oblitum (abbreviation n. o., plural nomina oblita) A nomen oblitum (Latin, forgotten name) is one that is technically a senior synonym of another, more recent name, but which has been used little or not at all since its original publication, and which would cause confusion if resurrected. Under the International Code of Zoological Nomenclature, to qualify as a nomen oblitum a name must not have been used as valid since 1899, and the competing junior name must have appeared in at least 25 works by at least 10 authors in the immediately preceeding 50 years and over a period not exceeding 10 years. The term "nomen oblitum" has also been used in the past for names suppressed by the International Commission on Zoological Nomenclature. A name that remains in place due to its senior synonym being a nomen oblitum is called a nomen protectum. (see ICZN online for more details)
For example, the name Tyrannosaurus rex Osborn, 1905 is a junior synonym of Manospondylus gigas Cope, 1892. However, because of the obscurity of the name Manospondylus compared to the name Tyrannosaurus, the former has been declared a nomen oblitum, and Tyrannosaurus rex remains the correct name.
Unlike the ICZN, the International Code of Botanical Nomenclature does not have any provisions for automatic rejection of an old name, requiring an action by the Commission for any name suppression. It is therefore not uncommon in botanical nomenclature for old names to be resurrected.
Outgroupin phylogenetics, a taxon that is not part of the clade under consideration, but is including in the analysis in order to provide a baseline. In cladograms, outgroups are shown branching off at the base of the tree.
Paleontologythe study of ancient life, on thebasis of fossil or other remains.
Pan-group, Total groupA crown group and its stem group considered together. The Pan-Aves thus contain the living birds and all (fossil) organisms more closely related to birds than to crocodiles (their closest living relatives). Pan-Mammalia are all mammals and their fossil ancestors down to the phylogenetic split from the remaining amniotes (the Sauropsida). Pan-Mammalia is thus an alternative name for the clade Synapsida. With the exception of a few taxa, such as turtles, the pan-group approach has not caught on because it results in unnecessary junior synonyms.
ParalectotypeIn taxonomy, all of the specimens in the syntype series of a species or subspecies other than the l lectotype.
ParatypeIn taxonomy, any specimens in the type series other than the holotype (or lectotype in the case of paralectotypes). Paratypes have no official status in determining species identity, but may have historical or practical significance (for instance, if the holotype does not show all the features useful in characterising the species). The term allotype is sometimes used for a paratype that represents the opposite sex from the holotype.
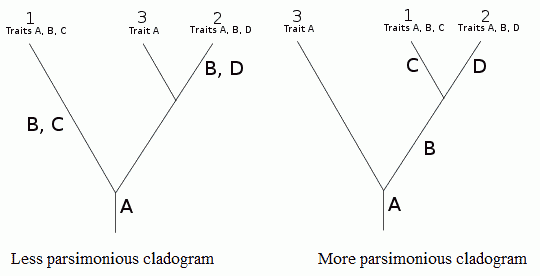 |
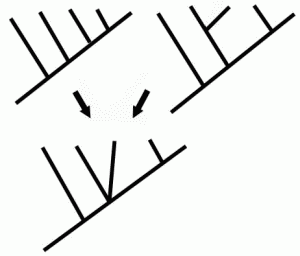
| Use of parsimony in cladistics. It is considered more likely that trait B evolved only once (right hand cladogram) rather than twice (left-hand cladogram).
|
ParsimonyAlso known as Occam's Razor (after the medieval theologian William of Ockham (c. 1285-1349), who rejected the idea of universals) is the principle that recommends when choosing between two competing hypotheses, that the simplest explanation of the evidence or observation is to be preferred, when the hypotheses are equal in other respects. A central premise in cladistics, where computer algorithms routinely generate huge numbers of cladistic trees. When reconstructing the phylogenetic relationships of a group of species or taxa, the principle of parsimony implies that we should prefer the branching pattern or phylogeny that requires the fewest number of evolutionary changes (see diagram at right), whether morphological, molecular, or both. Under maximum parsimony algorithms, the preferred phylogenetic tree is the one that requires the least number of evolutionary changes to explain the observed sets of characters (or traits).
The emphasis on parsimony dates back to the original hand-coded (pre-statistical algorithms) morphology-based cladistics of Hennig, and Hennigian paleontological cladists like Gauthier, Gaffney, and other early workers in the field who emphasised a small number of well-known synapomorphies as a way of constructing phylogenetic trees. Especially with molecular phylogeny, parsimony methods are particular vulnerable to long branch attraction. This is also the case with morphology-based phylogeny, when homoplasy, traits evolving at different rates, and phylogenetic incongruence come into the picture. It now seems that such factors are widespread if not endemic in the evolution of life, making dependence on parsiomony increasingly problematic. An example here is Archaeopteryx, which is resolved as a non-avian deinonychosaur using parsimony-based alogorithms, but as a true bird using maximum likelihood computation (see e.g. Nobu Tamura - Paleoexhibit). The frequent incongruency between morphological and molecular phylogenies is another example, consider for example the Afrotheria which make no morphological sense.
Pattern cladism, Transformed cladismDissenting Cladistic school, distinguished from phylogenetic or process cladism. Sometimes known as Cladists with a capital C (Williams and Ebach 2006). Transformed cladism is usually included here as well, although following Ebach et al 2008 they are given a separate entry. Founded by Gareth Nelson and Nelson Platnick ("New York Cladists") (Glossary of Phylogenetic Systematics - Günter Bechly although the latter is also associated with transformed cladism. (Ebach et al 2008). As with phenetics, character rooting and synapomorphies are not used, although monophyletic groups are acknowledged. Pattern Cladism asserts that a cladogram is merely a summary of shared characters, that could at best simply test a historical reconstruction (as a phylogenetic hypothesis), but reject the possibility that a real evolutionary history can ever be arrived at. Pattern cladistics no longer exists as an independent school, although its pragmatic empirical insights, such as cladistics as hypothesis testing, have been incorporated by mainstream cladistics.
Phenetics, Phenetic systematicsSchool of numerical taxonomy that classifies organisms on the basis of overall morphological or genetic similarity. It was abandoned in favour of cladistics for a number of reasons, including numerous difficulties encountered owing to convergence (homoplasy, as individual characters assumed to be homologous were not carefully analysed), mosaic evolution, and a shortage of diagnostic characters. With the rise of molecular systematics, distance methods, which are basically phenetic methods, have become popular, although these are vulnerable to the same problems, especially that of homoplasy.
Phenetic pattern analysisvery similar to Phenetics and generally synonymised, it uses numerical methods for taxonomic classification.
Phenetic Species Conceptphenetics-based definition that defines species as a set of organisms that look similar to each other andare distinct from other such sets (Ptacet & Hankison (2009)). Like phenetics, this is no longer used as it does not reference phylogeny
PhenotypeThe set of measurable or detectable physical or behavioral features of an individual. The phenotype represents the expression of the genotype of the individual as modified by environmental conditions during the individual's ontogeny.
 |
|
|
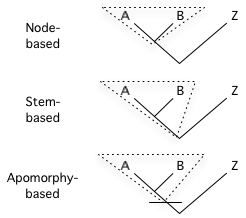
Phylocodeabbreviation for the International Code of Phylogenetic Nomenclature, a developing draft for a formal set of rules governing phylogenetic nomenclature. Its current version is specifically designed to regulate the naming of clades, leaving the governance of species names up to the rank-based codes. Unlike Linnaean-based nomenclatural codes the Phylocode does not require the use of ranks, although it does optionally allow their use. Rather than define taxa using a rank (such as genus, family, etc.) and a type specimen or type subtaxon, the content of taxa are delimited using a definition that is based on phylogenetic relationship and uses specifiers (e.g., species, specimens, apomorphies) to indicate actual organisms. The formula of the definition indicates an ancestor. The defined taxon, then, is that ancestor and all of its descendants. Thus, the content of a phylogenetically-defined taxon relies on a phylogenetic hypothesis. In the Phylocode, clades may be node-based, stem-based, or apomorphy-based (see diagram at right).
The theoretical foundation of the Phylocode was developed in a series of papers by de Queiroz and Gauthier, which was foreshadowed by earlier suggestions that a taxon name could be defined by reference to a part of a phylogenetic tree. The number of supporters for official adoption of the Phylocode is still small, and it is uncertain, as of 2011, whether the code will be implemented and if so, how widely it will be followed.
Phylogenetic hypothesisan empirical hypothesis regarding evolutionary relationships suggested through cladistic or other phylogenetic methods. Confusingly and despite the name, a phylogenetic hypothesis is not the same as phylogeny, does not purport to describe the actual course of evolution itself, complete with ancestor descendent relationships. Rather it is a stylised or abstract representation of this (usually in the form of a cladogram or similar), based on available data, with the proviso that this can and indeed is likely to change or even be radically revised with new data, discoveries, and analyses.
Phylogenetic incongruencewhen two equally persuasive, verified, robust, and empirically supported methodologies give contrary phylogenetic results. For example, using morphology, the soft shelled turtles (Trionychia) are the most derived group of cryptodires, whereas using molecules, they are the most basal group. The problem here is in deciding which, if any, of the two methodologies provides the more reliable phylogenetic signal
Phylogenetic nomenclature (or classification, or taxonomy)classification and taxonomy based on cladistic (Phylogenetic systematic) principles ("vertical" ancestry, not "horizontal" similarity), proposed as a rank-free alternative to the Linnaean system of classification, redefining taxa previously named under evolutionary systematics (e.g. Synapsida), and accepting only monophyletic clades. The goal is to make classification synonymous with phylogeny; i.e. to get rid of similarity altogether. Phylogenetic nomenclature has led to a number of controversial proposals, such as the abandonment of Linnaean binomial nomenclature, the rejection Linnaean ranks, and the migration of established names to crown clades (Benton 2007, p.651); e.g. Tetrapoda (this last reflecting an emphasis on neontology over paleontology that is still found in cladistics). Despite the logical and theoretical appeal of this approach, there are still problems in applying it in practice (Carlson, 2001, p.1113). See also Phylocode.
Phylogenetic signalthe amount of information, or "signal" that can be retrieved from the background "noise" of any phylogenetic analysis. It is only to be expected that the advocates of any particular methodological paradigm consider that their own methodology provides the clearest phylogenetic signal. Therefore, in the case of any phylogenetic incongruency between themselves and a rival methodology, their own paradigm is automatically to be preferred. Take the example of molecular phylogeny verses cladistic morphology. Morphology strongly supports a monophyletic Insectivora, based on a large number of unique shared characteristics, whereas molecular sequencing indicates homoplasy and divides the insectivores into two unrelated clades, placing them in groups for which there is no morphological support. Because molecular phylogeny has replaced cladistics as the default option for any analysis that includes extant (recent) taxa, unqualified support of phylogenies resulting from this methodology, despite still being problematic are the standard approach. The unspoken implication here is that molecular phylogney has a much higher and more reliable phylogenetic signal, and that morphology involves so many convergences and reversals as to make extracting any possible phylogfenetic signal almost impossible, without first being grounded in the molecular tree. Here at Palaeos we have tried to adopt a non-partisan approach incorporating all methodolgies, popular and unpopular, the only proviso being that be scientific, verifable, and found in earlier or recent scientific literature.
Phylogenetic systematicsAlso known as Hennigian systematics, Numerical cladism, Phylogenetic cladism, and Process cladism. Cladistic methodology that derives from Hennig's work and that of others such as James S. Farris, Walter Fitchand, and Herb Wagner. States that only shared derived characters can provide information about phylogeny. Those taxa that share a greater number of shared features are considered more closely related than those that don't. However, the shared characteristics have to be advanced (derived) rather than on primitive. The relationship between them is shown in a branching hierarchical tree called a cladogram. The cladogram is based on the principle that the fewest number of changes to map all the changes of character states is the most likely one; called the principle of parsimony. Only monophyletic groups are recognised. Unlike pattern cladism , which only aims at the calculation of most parsimonious cladograms from large data-sets, phylogenetic systematics also seeks to reconstruct phylogenetic schemes, in which all branching points are convincingly supported by characters, and using optimization (transformation series) (sensu Farris 1983) to select from a number of possible trees.
Phylogenetic tree See Tree.
PhylogeneticsA term derived from the Greek φῦλον, phulon, "tribe", γενέτης, genetēs, "ancestor" and -ικός, -ikos, an adjective-forming suffix. (Perseus Digital Library, Wiktionary) The synthesis of cladistics and molecular phylogeny, phylogenetics is the study of evolutionary relatedness among groups of organisms (e.g. species, populations). It analyses molecular sequencing and morphological data matrices, using statistical methods such as maximum parsimony, bayesian inference and maximum likelihood. This phylogenetic analysis is used to determine the most likely phylogeny that would correspond to an actual tree (phylogram or cladogram) of a particular shape. The tree represents the evolutionary history of a group. Phylogenetics has also begun to incorporate other fields such as evo-devo) which implies this is all leading to a new evolutionary synthesis (replacing the 20th modern synthesis). See also Daniel F. Simola - Molecular Evolution and Phylogeny (pdf) for synoptic overview.
Phylogenomicscatch-all label for the intersection between the fields of evolution and genomics. The use of cladistic principles to interpret genome data, and better understanding of gene function. One branch of phylogenomics involves the use of these data to reconstruct the evolutionary history of organisms. It considers molecular data from many genes, or even whole-genome approaches, rather than just a few specific genes, and using broad taxon sampling. Such studies have provided insights into the relationships of protostome phyla that were previously obscure, and allowed resolution of long-standing questions such as the relationships of Arthropoda and Onychophora and various trochozoan phyla. Another application, "Pharmacophylogenomics" is the use of phylogenomics in aid of drug discovery, through improved target selection and validation.
PhylogenyA distrinction can be made between trhe scioence of phylogeny and phylogeny itself. The term was coined by Haeckel (Haeckel 1866) to refer to the science of the study of the family history of life, the evolutionary relationships among groups of organisms, often illustrated with a branching diagram called a tree. Phylogeny in itself therefore refers to the evolutionary history of a group through deep time; in other words, the evolutrionary tree of life. There are several different forms:
Haeckelian/evolutionary systematics maps the inferred lines of descent of a group of organisms, in order to reconstruct the common ancestors of that group, map the amount of divergence among the descendants of the common ancestor, and explore the evolutionary history of a group of organisms (Mayr & Bock 2002 p.192). Evolutionary Systematics, which is highly phylo-optimistic, seeks to reconstruct the actual evolutionary history of a group, including the actual common ancestor (either an ancestral species or, more usually, supra-specific taxa), and, with the help of the fossil record, tracing the evolution from ancestor to descendants and from there to further descendants.
Hennigian cladistics or phylogenetic systematics (Hennig (1950, 1966)) is the analysis of and relationship between monophyly taxa (clades), using shared unique characteristics (synapomorphies). Despite the name, phylogenetic systematics does not describe the actual evolutionary history of life, but rather the construction of phylogenetic hypotheses, represented graphically in the form of cladograms. It rejects the idea of an actual common ancestor a la Evolutionary Systematics, and instead posits a hypothetical common ancestor
Molecular phylogeny is the use of molecular data (derived for example from DNA, RNA, and/or protein sequencing) for phylogenetic analyses. Generally speaking, the more closely related two organisms are, the more similar their gene sequences will be. By comparing the similarities and differences we can deduce the pattern of how those organisms are related. Because of the emphasis on empiric data it is less convcerned with testing rival hypotheses than cladistics is. However it shares with modern statsitical-based cladistics the emphasis on computer algoritrhms and a bifurcating tree of life.
Other approaches are possible too, for example developmental. In this way, phylogeny is used to understand the evolutionary history of life on Earth.
Phylogeographyresearch field that investigates the principles and processes that govern the geographic distributions of genealogical lineages, especially those within and among closely related species.
PhylogramA phylogenetic, molecular-based dichotomous branching tree that resembles a cladogram, although it differs in that the branch lengths are proportional to the amount of inferred (quantitatively measured) evolutionary change. Phylograms therefore convey more information than cladograms. Whereas cladograms only give information about the branching order and nothing else, phylograms also include information on the amount of change. Unlike chronograms they do not include stratigraphic range (unless one could draw them in three dimensions, like a hologram, perhaps), in a rooted phylogram evolutionary divergence takes the place of the time axis in a cladistic chronogram.
PhylumIn the Linnaean classification the taxonomic rank between kingdom and class, and hence one of the highest levels of taxonomic classification, used to define major groups of organisms; e.g. molluscs, arthropods, echinoderms, chordates. Phyla can be thought of as groupings of animals based on a shared general body plan. What this means is that despite the seemingly different external appearances of organisms, they can be classified into phyla based on their internal and developmental organizations. Despite their obvious differences, spiders and barnacles both belong to the phylum Arthropoda; but earthworms and tapeworms, although similar in shape, belong to different phyla. Although Linnaean rankings are not used in cladistic analysis, the majority of phyla are still accepted as they constitute monophyletic clades. (There are a few exceptions; e.g. growing consensus on the basis of molecular phylogeny is that Porifera (sponges) constitute an evolutionary grade. The rank of Phylum was not in Linneaus' original classification system, but was coined later by Haeckel.
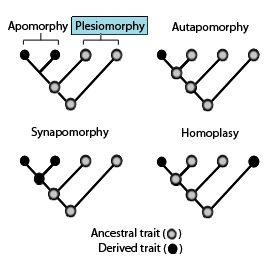 |
| Plesiomorphy in relation to apomorphy, autapomorphy, synapomorphy, and homoplasy.
|
Features shared more widely than in a group of interest. These are primitive for the group in question and cannot provide evidence for the group. An evolutionary trait that is homologous within a particular group of organisms but is not unique to members of that group (compare apomorphy) and therefore cannot be used as a diagnostic or defining character for the group. For example, vertebrae are found in zebras, cheetahs, and orangutans, but the common ancestor in which this trait first evolved is so distant that the trait is shared by many other animals. Therefore, possession of vertebrae sheds no light on the phylogenetic relations of these three species.
A Dictionary of Biology, Oxford University Press, © Market House Books Ltd 2000
Polarityin phylogenetic cladistics this refers to the ordering of a particular character state, determined either independently of tree construction (direct method) or more usually from a rooted tree (indirect method) (Michael D. Crisp - Introductory glossary of cladistic terms) To quote Telford & Budd 2003 p.487: "In order for an analysis to be useful in an evolutionary sense, it needs to be rooted, in other words we need to know the polarity of change of the characters that interest us. If we consider two taxa in isolation (say a lizard and a mouse) that differ in a certain character (e.g. hairless or hairy) how do we know which of the two has the primitive character state and which the derived? ...(T)o determine the direction in which the evolution of this character has proceeded...knowledge of the state of the character in a species that is an outgroup (is needed)...in this case, a frog would be appropriate. As the frog is hairless, parsimony suggests that hairlessness is the primitive character and we can infer from this that hair has evolved in the lineage leading to mice after this lineage had diverged from reptiles." Polarity is one of the ways in which phylogenetic systematics is distinguished from non-phylogenetic ordering systems such as phenetics and pattern cladistics.
Polyphyly, Polyphyletic groupA group that does not share a common ancestor, but is defined on the basis of independently acquired or convergent (non-homologous) character states. Examples for polyphyletic groups would be the old taxon Pachydermata which includes the thick-skinned hippos, rhinos and elephants, or the taxon Haemothermia (endorsed at one time by Lovtrup and Gardiner) for a grouping of haemothermic birds and mammals. Polyphyletic groups are considered invalid by both evolutionary and phylogenetic systematics.
Polytomyin a cladistic phylogeny, a node where more than two lineages descend from a single ancestral lineage. This indicates either that we don't know how the descendant lineages are related or the descendant lineages speciated simultaneously. Where a branching pattern cannot be resolved, the branches in question can be collapsed to show the absence of a hypothesis for the relationships among the lineages that they represent.
Rankthe hierarchical level of a supra-specific taxon, according to the Linnaean approach to classification. The eight ranks are kingdom, phylum (added by Haeckel), class, order, family, tribe (used mostly in botany, much more rarely in zoology and paleontology), genus, and species, plus optional intermediate grades represented by the suffixes super-, sub- and infra-. ( In the three domain theory of Carl Woese and co-workers, a further rank, domain, is sometimes added above kingdom, although it seems to me that domain and kingdom are just different ways of approaching the same topic (like evolutionary and phylogenetic systematics)). The highest ranks are the most general, whilst each sub-division or rank adds it own increasingly specific and unique characteristics. In this way any organism or species can be grouped more and more specifically within the hierarchy. A central part of evolutionary systematics, according to which every taxon that evolves from another taxon has the same taxonomic rank. So class Reptilia would give rise to class Mammalia, not subclass Mammalia (although, Reptilia includes a number of subclasses, these are still part of the class Reptilia). Determining the appropriate rank is an art, not a mechanical process, and inevitably ranks don't always equate. e.g. the orders of modern birds are probably equivalent to families or superfamilies of fish or invertebrates. Ranks are strenuously rejected by most cladists, although some paleontologists such as Michael Benton argues that cladistics and Linnaean ranks are not be incompatible. Moreover, ranks are useful in everything from field guides (compare the muddled organisation of taxa in Greg Paul's otherwise superlative Princeton Field Guide to Dinosaurs with the clear arrangement of Predatory Dinosaurs of the World from more than two decades earlier) to measuring the degree of biodiversity through time.
Replacement nameA name that is assigned to replace a name that is a junior homonym, e.g., Onceropyga Turner, 1904 is the valid name and Onceropyga Ferris, 1955 is the junior homonym and must be replaced; Hoy (1963) proposed the replacement name Oregmopyga.
Scala Naturaea Latin expression meaning "natural ladder", is a sort of proto-taxonomy first developed by Aristotle, according to which the natural world can be arranged in a single linear series from inanimate matter through plants, invertebrates, higher vertebrates, and finally man. Along with Plato's Principle of Plenitude it led to the idea of the Great chain of being. Scala Naturae and Great Chain of Being remained central ideas in natural philosophy until the mid 19th century.
Semistrict consensusalso called "combinable component" consensus. If a particular grouping in one tree is not contradicted by the other trees, it will be retained in the consensus. When there is a conflict in grouping, semistrict consensus behaves like strict consensus.
Senior homonymIn taxonomy, the oldest described homonym, e.g., Onceropyga Turner, 1904 is the senior homonym and Onceropyga Ferris, 1955 is the junior homonym.
Senior synonymIn taxonomy, the oldest synonym, e.g., Apiomorpha pharetrata Scharder, 1863 is the senior synonym and A. nux Fuller, 1896 is the junior synonym.
Sequencingany of several methods and technologies that are used for determining the order of proteins in a cell, or nucleotide bases (adenine, guanine, cytosine, and thymine) in a molecule of RNA or DNA. An essential element in modern biological systematics (molecular phylogeny). The rapid speed of sequencing attained with modern DNA sequencing technology has been instrumental in the sequencing of the human genome (the Human Genome Project). Related projects, often by scientific collaboration across continents, have generated the complete sequences of many animal, plant, and microbial genomes.
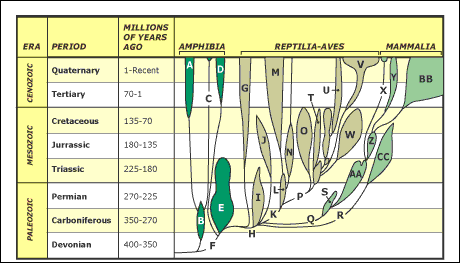 |
| The spindle diagram shown here is typical of mid 20th century phylogenies. This particular diagram which I found through Google image search is from an online creationist book ( original url). The taxa correspond to orders, subclasses, or classes, in the Linnaean ranking. The caption reads: Fig. 20. Family tree of the vertebrates. On the left is a geological time scale. Phylogenetic origins of various groups of vertebrates. A) Urodela, B) Lepospondyli, C) Apoda, D) Anura, E) Labyrinthodontia, F) Apsidospondyli, G) Chelonia, H) Anapsida, I) Cotylosauria, J) Euryapsida, K) Diapsida, L) Eosuchia, M) Squamata, N) Rhynchocephalia, O) Ornithischia, P)Thecodontia, Q) Synapsida, R) Parapsida, S) Pelycosauria, T) Pterosauria, U) Crocodilia, V) Aves, W) Saurischia, X) Prototheria, Y) Metatheria, Z) Pantotheria, AA) Therapsida, BB) Eutheria, CC) Ichthyosauria.
The width of the spindle shows taxonomic diversity, numeric abundance (the abundance of fossils of the group in strata of the particular geologic period), or both; the distinction here being often poorly defined. In an effort to introduce precision, spindle width may be drawn according to the number of genera or families known from a particular time period, but even determining what qualifies as a genus or family can be arbitrary (see splitters versus lumpers). Contrast this diagram with cladistic dendrograms and cladograms which show relationships between individual species, without referencing time, transformation, or evolutionary lineage.
|
Spindle diagramA evolutionary tree that maps lineage diversity or abundance mapped against geologic time. They are called spindle diagrams because each lineage generally begins at a point, widens in the middle (representing increasing diversity over the course of millions of years), and then declines towards the top (representing the dwindling fortunes of the lineage in question). They are similar to modern chronograms (or clado-chronograms) except that they convey additional information in the form of the diversity and/or abundance of each lineage at a particular time (as represented by the width of each "spindle". Spindle diagrams are employed in evolutionary taxonomy. They provide a purported map of actual phylogeny rather than a hypothesis, as they emphasise ancestral or paraphyletic groups, transitional forms, and the transformation of one group into another, showing where more recent lineages emerge from earlier ones. Also called a Bubble diagram or Romerogram. Cladistic formalism rejects the use of spindle diagrams. This doesn't mean that spindle diagrams are invalid or untrue, only that cladistics and phylogenetics speak a different language.
Splitters versus lumpers Philosophical conflict among taxonomists, as regards ranking of a taxon. As the name indicates, splitters tend to divide varying individuals from a single species among several different species. Lumpers tend to include specimens or populations normally attributed to different species in a single species. The same principle can be applied at the supra-specific level. (MAK111018)
The earliest use of these terms was apparently by Charles Darwin himself, in a letter to J. D. Hooker in 1857.
Those who make many species are the 'splitters,' and those who make few are the 'lumpers.'
They were introduced more widely by the biologist George G. Simpson in his 1945 work The Principles of Classification and a Classification of Mammals. As he put it:
[S]plitters make very small units — their critics say that if they can tell two animals apart, they place them in different genera (...) and if they cannot tell them apart, they place them in different species. Lumpers make large units — their critics say that if a carnivore is neither a dog nor a bear, they call it a cat.
Stem groupNot to be confused with stem-based group (see above entry), the concept of stem group is used in phylogenetics to cover extinct evolutionary "aunts" and "cousins" of living groups. A crown group is a group of closely-related living animals plus their last common ancestor plus all its descendants. A stem group is a set of offshoots from the lineage at a point earlier than the last common ancestor of the crown group; it is a relative concept, for example tardigrades are living animals which form a crown group in their own right, but Budd 1996 and 2001 regarded them also as being a stem group relative to the arthropods. Stem group shown in yellow in this diagram. (Wikipedia). The distinction however between the stem- and crown group is an arbitrary one, because it is determined only by the most basal member of the crown group that is still extant; if Branchiostoma had gone extinct in the Pleistocene, or even the 15th century, our concept of the euchordate crown-group be radically different, because Branchiostoma lacks many features of higher chordates (Budd 2001 p.265).
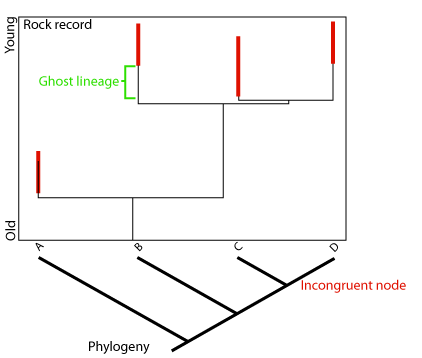 |
| How do we identify ghost lineages and measure their prevalence in a cladogram. All other things being equal, we expect the terminal taxa that branch off of a cladogram first to appear first in the fossil record. When this is true, the cladogram is said to be stratigraphically congruent. Often, cladograms are not stratigraphically congruent. This happens when there are long ghost lineages.
|
 |
| Two types of phylogeny: star and hierarchical
|
Stratigraphic congruencealso called stratigraphic consistency, the degree to which the terminal taxa that branch off of a cladogram match the order with which they first appear in the fossil record. A simple measure of stratigraphic congruence is the Stratigraphic Congruence Index (SCI) of Huelsenbeck (1994) is defined as the proportion of stratigraphically consistent nodes on the cladogram, and varies from 0 to 1. A node in the cladogram is consistent with trhw fossil record when the first occurrence of the taxa above it (daughter clades etc) are younger or equal in age to those below it (parent clades etc). Also when it is the same age or than the first occurrence of its sister taxon.
Stratigraphyin the phylogenetic context, the chronological and stratigraphic order by which taxa appear in the fossil record, in this context refers to biostratigraphy, rather than stratigraphy as such. Here, the more primitive or ancestral form should always preceed the more advanced type of organism. So, for example, the protodinosaurs ("non-dinosaurian dinosauromorpha" to use the unwieldly cladistic definition) appear in the middle Triassic period, whereas their descendents the true dinosaurs only appear some ten or twenty million years later in the late Triassic. However the vagaries of the fossil record mean that not every evolutionary lineage is recorded, for this reason most cladists, apart from the rarely used stratocladistic approach, ignore stratigraphic sequence and fill in the gaps with ghost lineages.
Stratopheneticsphylogenetic method based on (a) the identification of taxa based on phenetic similarities among specimens and (b) stratigraphic interval, so that taxa from different time-intervals are linked in presumed ancestor-descendant sequences according to their similarities.
Strict consensusthis is the most conservative consensus method used in in cladistic analysis, which only recognises clades that appear in all of the trees. It's advantage is that it only includes data that is totally unambiguous. The disadvantage is that it is thrown off by the slightest difference. For example, two trees may be identical except for the placement of a single sequence, yet their strict consensus tree might be completely unresolved. The resulting is a "star" phylogeny, a broad polytomy with only radiating lines, and very little or no resolution or phylogenetic structure. This is shown by the blue cladogram on theright, which is placed next to a more conventional, branching phylogeny.
SubgenusA group of species less inclusive than a genus. The subgenus name is written in italics and brackets, after the genus but before the species. It may be the same as or different to the genus, e.g. Cypraea (Cypraea) tigris Linnaeus, the tiger cowrie, belongs to the subgenus Cypraea of the genus Cypraea. However, it is not mandatory, or even customary, when giving the name of a species, to include the subgeneric name. One paleo artist and author of popular books on dinosaurology, Greg Paul, sometimes coins subgenera, although this practice is otherwise very rarely used in vertebrate paleontology
SubspeciesThe smallest taxonmic rank; a group of organisms less inclusive than a species. The term is usually applied to populations or groups within a species that have distinct forms or characteristics and live in a restricted geographic area. In contrast to the species, members of different subspecies can usually interbreed and give rise to fertile offspring. The subspecies name is written in italics after the species, and may or may not be the same as the species name. e.g. The Cape Mountain Zebra is referred to as Equus zebra zebra, as distinguished from Hartmann's Mountain Zebra, Equus zebra hartmannae
SupermatrixOne of the new developments in cladistics that have become possible through cheap and powerful computing, supermatrixes involve simultaneous analysis of all available character data. Rather than separate analyses of data sets and subsequent integration of the resulting trees (supertree), all character data is considered simultaneously to enable incorporation of diverse kinds of data, including characters from fossils, morphology, and molecular phylogeny.
 |
| Dinosaur supertree
|
SupertreeIn cladistics, a "supertree" refers to the synthesis of a number of distinct cladograms, combining morphological, molecular, and other data from the different individual phylogenies. Supertrees result from combining many smaller, overlapping phylogenetic trees into a single, more comprehensive tree. They are distinguished from classic consensus techniques in that the source trees need only have overlapping rather than identical taxon sets. Because supertree construction uses other tree topologies rather than the primary data underlying those trees, they can be constructed using all available phylogenetic hypotheses, even those based on incompatible data types, or lacking data entirely. Supertree have produced phylogenies of a number of large taxonomic groups. However supertree strength is also its weakness, and this approach has been harshly criticised by systematists precisely because it only considers the topology of the source trees, effectively discarding primary data. A supertree or quasi-supertree approach is also standard with ASCII phylogenetic trees. Supertree construction is probably as old as the field of systematics itself, and remains our only way of visualizing the Tree of Life as a whole.
Symplesiomorphy In cladistics, a shared plesiomorphic character trait, which is shared between two or more taxa, but which is also shared with other taxa which have an earlier last common ancestor with the taxa under consideration. An example is pharyngeal gill breathing in bony and cartilaginous fishes. The former are more closely related to Tetrapoda (terrestrial vertebrates, which evolved out of a clade of bony fishes) that breathe via their skin or lungs, rather than to the sharks, rays, etc. Their kind of gill respiration is shared by the "fishes" because it was present in their common ancestor and lost in the other living vertebrates. Contrast with apomorphy/synapomorphy.
SynapomorphyIn cladistics, an apomorphy that is shared by (syn-) by several taxa, where the trait in question originates in their last common ancestor. Being shared by multiple taxa, synapomorphies can be used to diagnose (describe) a clade (a monophyletic group). Compare with homology. True synapomorphies usually are a given set of terminal groups, shared by two or more terminal taxa, but this is not essential to the concept. Thus, if some descendants of a last common ancestor possess a synapomorphic trait, in the case of reversals it is not strictly necessary that all of its descendants must possess the same trait. Contrast with plesiomorphy and homoplasy, which are shared primitive and shared convergent characteristics also of no phylogenetic value.
Synonym. In taxonomy, the term synonyms is used to refer to two or more names referring to the same taxonomic entity. It is a general principle of taxonomy that any taxon can have only one valid name - usually, this is the oldest name available (the senior synonym, as opposed to a junior synonym) if there is more than one (but see Nomen oblitum for one example of where this rule may be suspended). A list of names used to refer to a taxonomic entity is referred to as a synonymy.
Synonyms may be either objective or subjective. Objective synonyms have the same type as each other, and as such will always refer to the same taxon. Subjective synonyms have different types, and authors may differ as to whether they represent the same taxon or not. In synonymies presented on Palaeos.org, we have generally distinguished between the two classes by using "=" for objective synonyms and "incl." for subjective synonyms.
SynonymyA section of a systematic presentation about an organism that lists all of the names that have been used for the organism including synonyms, new combinations, misidentifications, etc. In some cases this section may include only true synonyms.
SyntypeThe series of specimens used to describe a species or subspecies when the author did not include a holotype ( ScaleNet - Terms Pertaining to Zoological Nomenclature). Where the original description was based on a number of specimens, some or all of them may hold equal status as type specimens. Should a syntype series turn out to contain examples of more than one species, a subsequent reviser may designate a lectotype.
Systematic biology, taxonomy, and scientific classification are often confused and used interchangeably. However, taxonomy is more specifically the identification, description, and naming (i.e. nomenclature) of organisms, classification focuses on placing organisms within hierarchical groups that show their relationships to other organisms, and systematics alone deals specifically with relationships through time, and can be synonymous with phylogenetics, broadly dealing with the inferred evolutionary hierarchy of organisms.
Taxonomic inflationPejorative term for what is perceived to be an excessive increase in the number of recognised taxa in a given context, due not to the discovery of new taxa but rather to putatively arbitrary changes to how taxa are delineated. For example, a subspecies may be elevated to species rank, through the arbitrary decision that the differences between the various taxa warrant distinguishing them at species rank. (From Wikipedia). Another form of taxonomic, or rank, inflation is elevating subfamilies to families, families to superfamilies or orders, and so on, which tends to be an on-going process as more taxa are discoevred. For eaxmple in the late 1980s Carcharodontosaur theropods were included under the Allosauridae, now they are given their own family.
TaxonomyThe field of science converned with discovering, describing, clasisfying, and naming organisms. It is supported by institutions holding collections of these organisms, with relevant data, carefully curated: such as Natural History Museums, Herbaria and Botanical Gardens. Taxonomy uses taxonomic units, known as taxa (singular taxon). In addition, the word is also used as a count noun: a taxonomy, or taxonomic scheme, is a particular classification ("the taxonomy of ..."), arranged in a hierarchical structure. (Wikipedia) The roots of taxonomy go back to Aristotle at least, although it was only really developed as a modern science by Linnaeus. Modern approaches to taxonomy follow the same principle of organising and understanding the natural world. These taxonomies fall into three major schools: phenetic, phylogenetic (cladistic), and evolutionary. Each of these pertains to a different phylogenetic methodology, a different way of mapping out the history and evolution of life on Earth . . In biological taxonomy there are, according to Ereshefsky (2000, p. 7) "no fewer than four general schools of taxonomy: evolutionary taxonomy, pheneticism, process cladism, and pattern cladism". Each of those schools have their own view on how to get from the characteristics of an individual organism to a species, and also the meaning of the term species varies between schools of taxonomy. (cited from Birger Hjørland) See also Alpha taxonomy , cladistics, Linnaean classification, Systematics.
Tetrapodfour-legged, land-living vertebrate, or any secondarily limbless (e.g. snakes) or aquatic (e.g. whales) descendants of such. Cladistic terminology disagrees over whether "tetrapod" should be used to include all four-legged animals (stem-based definition) or only those that include the common ancestor of all living tetrapods and its descendants (crown-based definition).
Terminal, Terminal taxonNot the end of the evolutionary line, but in cladistic formalism, one of the units whose collective phylogeny is reconstructed; shown diagramatically as the undivided tips of a cladogram. Terminals may be higher taxa, species, populations, individuals, fossils or even genes. There should be some rational basis for accepting the integrity of each terminal (for the purpose of the analysis), e.g. a monophyletic or diagnosable unit. Despite the claims by some authors, terminals do not need to be monophyletic; in fact, many species-level terminals are unavoidably paraphyletic. However, higher taxa used as terminals should be monophyletic.
Three-domain systembiological classification introduced by Carl Woese that rejects the old prokaryote-eukaryote distinction and divides cellular life forms into Archaea, Bacteria, and Eukarya domains (usually interpreted as a taxonomic grade above kingdom). Woese argued that, on the basis of differences in 16S rRNA genes, the three groups each arose separately from an ancestor with poorly developed genetic machinery, called a progenote. To reflect these primary lines of descent, he treated each as a domain, divided into several different kingdoms. He conjectured an era in which there was a considerable amount of lateral transfer of genes between organisms. Species formed when organisms stopped treating genes from other organisms with equal importance to their own genes. Lateral transfer during this period was responsible for the fast early evolution of complex biological structures.
TopotypeOne or more specimens collected at the same location as the type series regardless of whether they are part of the type series.
Triply paraphyletic groupa group or taxon that is paraphyletic because three of its descendant lineages are not included. There would also be quadruply paraphyletic groups and so on. See also singly and doubly paraphyletic groups
TypeThe term "type" is tied in biological nomenclature to a very specific concept - that of a designated specifier that provides the definitive concept of a given taxon. For instance, when describing a new species, the author(s) is required to name the specimen or one of the specimens used as the type specimen. The name for the new species then becomes indelibly tied to that specimen, and should any confusion ever arise as to the identity of the species (for instance, if it turns out that two or more species have been mistaken for one, or if the published description turns out to omit some feature[s] required for identification), examination of the type specimen should (hopefully) resolve these issues. Similarly, at higher levels, each genus requires a type species, and each family requires a type genus. See International Code of Zoological Nomenclature Online for more information relevant to animals, and International Code of Botanical Nomenclature for plants. Different nomenclatorial codes may differ in the terminology used.
A number of terms are in use to refer to different classes of types, including holotype, (the most important, as it used to define a species), allotype, epitype:, hapantotype, isotype, lectotype, neotype, paratype , syntype, topotype, type series and type strain.
Type genusA genus that has been selected as the standard bearer of a tribe, family, or superfamily and provides the stem of the family-group name.
Type localityThe geographic location where the primary type was collected.
Type seriesthe total group of specimens used in the original description. Ideally, one specimen is the holotype and the remainder paratypes, but if no holotype has been designated, the entire type series become syntypes.
Type speciesA species that has been selected as the standard bearer of a genus or subgenus.
Type strain (Bacteriological Code) For prokaryotes, the type is not a preserved specimen, but an isolated culture. The Bacteriological Code of Nomenclature requires that cultures of the type strain be deposited in at least two separate institutes' culture stores.
Unavailable namein taxonomy, a name that is incorrectly proposed according to the International Code of Zoological Nomenclature.
Unjustified emendationin taxonomy, an emendation that is incorrect according to the International Code of Zoological Nomenclature, e.g., the generic name Hemiberlesea Lindinger is an incorrect change of Hemiberlesia Cockerell according to the Code and is an unjustified emendation.
Valid nameThe correct name of an organism, e.g., if Apiomorpha nux Fuller, 1896 and A. pharetrata Scharder, 1863 apply to the same species (and therefore are synonyms), then by the law of priority (the oldest name prevails) A. pharetrata Scharder, 1863 is the valid name.
Vraagteken effectfrom the Dutch "question mark", a term introduced by Schram and Hof (1998) to the effect that the absence of critical information has in destabilising cladograms. They found that by introducing fossil taxa, for which, obviously, many character states are unknown, and hence coded as a question mark in the matrix, resulted in a great variation in the topology of the trees recovered by parsimony analysis. Cladistic algorithms respond to the ambiguity caused by the missing data by generating a large number of equally parsimonious phylogenies .
Wastebasket taxona taxon that includes all species or groups that cannot be easily or conveniently placed elsewhere, e.g., for a while all large theropod dinosaurs that could not be included under the Ceratosauridae, Allosauridae or Tyrannosauridae were named Megalosaurus.
Weightingin cladistics, the empirically controversial (because non-quantifiable) yet necessary task of determining the phylogenetic significance of a particular character trait. For example, if there are three species of animals, one with brown fur, another with black fur, and one with brown scales, the presence or absence of fur is more important than the external colour, and hence would be given greater weight in phylogenetic analysis. Weighting is unavoidable if one is to address the problem of homoplasy vs homology.
Zombie taxon, Zombie effectBefore the zombie craze took over geek/nerd culture (perhaps as a counterpole to the excessively feminine/romantic "supernatural romance" vampire story) the technical term for a fossil of this sort was term "reworked". Refers to a fossil such as a dinosaur tooth that was washed out of sediments and re-deposited in rocks and/or sediments millions of years younger. This basic mistake in the interpretation of the age of the fossil leads to its title. The discovered fossil was at some point mobile (or "walking") while the original animal or plant had long been dead.
A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z
content by MAK110419, edited RFVS111203, last modified MAK120326, merged with taxonomy glossary 130316














