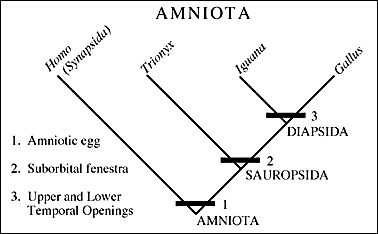
Cladogram by Paul Olsen (original url), showing four species (human, turtle, lizard and bird), according to their synapomorphies (shared unique characteristics). Most computer-generated cladograms involve many hundreds of characteristics
| Cladistics | ||
| Systematics | Cladistics Home |
| Systematics | ||
| Evolutionary systematics | Molecular phylogeny |
 Cladogram by Paul Olsen (original url), showing four species (human, turtle, lizard and bird), according to their synapomorphies (shared unique characteristics). Most computer-generated cladograms involve many hundreds of characteristics |
Cladistics is a rigorous methodology first developed by German entomologist Willi Hennig (who used the term "Phylogenetic Systematics"). It is based on three principles:
Cladistics acknowledges only Monophyletic groupings as valid. Paraphyletic groups (accepted in Evolutionary Systematics) and Polyphyletic groups are rejected as invalid, as is the whole Linnaean hierarchy above species rank (although sometimes taxa such as family etc are used in a more limited context).
The cladistic revolution of the 1970s and 1980s constituted a major paradigm shift in biology and systematics, with the Evolutionary system falling out of favour and being replaced by the one. Cladistics is based not on morphological similarity (as in the Linnaean system and more recently phenetics) or on ancestor and descent relationship (as in Evolutionary systematics) but in sister-group relationships between related taxa. Although originally based on recent organisms (neontology) it also can be used to analyse fossils. Indeed, when computer-based cladistic analysis came into its own in the 1990s, paleontologists were among the first zoologists to almost wholeheartedly adopt the system (Brochu & Sumrall, 2001).
Although the relation between cladistics and evolutionary systematics could be described as the difference between "Vertical" and "Horizontal" Taxonomy, the two systems are quite distinct and to some degree incompatible. This does not mean that one is right and the other wrong.
Contrary to popular belief, cladistics does not describe the actual evolutionary path of life. That is, it is not concerned with or describe the evolution of later organisms from common ancestors in the way that, say, Darwin or more recently Richard Dawkins do, and what the Evolutionary systematics of Romer and Simpson also describes. It simply provides a means of determining in which way (i.e. the branching order) living organisms are related to each other. Cladograms, in other words, are not evolutionary trees. What cladistics does do is provide a more precise and verifiable method of creating and testing phylogenetic hypotheses regarding the evolutionary relationships of past and current organisms. In this way, cladistic methodology can even be used to predict properties of yet-to-be discovered organisms.
The current trend in evolutionary thinking is to use statistical-cladistic methods to combine morphological and molecular data in large phylogenetic trees. When there is a conflict between the two methodologies, the tree derived from molecular phylogeny is most often preferred, although there is no empirical reason why molecular sequencing should be preferred over morphological studies, as both are equally robust. The Total Evidence approach provides a more balanced assesment by (ideally) giving equal weight to both methodologies. MAK130324
A few random links: Phylogenetics Primer - Douglas Theobald, (recommended), Introduction to Cladistics - UCMP, also really good, Am I a pattern or transformed cladist? (mail list anecdote on phenetics and cladists); Peter Forey - Cladistics for Palaeontologists (pretty technical). MAK111014
page MAK111014, edited RFVS111203, last modified MAK130324