|
|
Pieces |
| Fungi |
The Cell Wall |
The Cell Wall
A Spoonful of Sugars
Since we haven't done this elsewhere, it's time we provided the rudiments of sugar saccharide) chemistry, so that we can make useful noises about polysaccharides sugar polymers) -- easily the most common class of biopolymers on the planet. A more extensive and far better introduction may be found at Natural Products.
All sugar monomers of biological importance have structural formulas which looks something like this: CH2OH-(CHOH)n-CHO. In other words, they consist of a chain of carbon atoms, in which each carbon atom has a hydroxyl (-OH) group attached to it, except for C1 (sometimes C2) which has an aldehyde or keto (=O) group. In living organisms, the chain is generally 3-7 carbons long. In biologically important polysaccharides, the monomers are almost always 5- or 6-carbon sugars.
We have only reluctantly provided a reference graphic of a sugar monomer in linear form because, in life, 5- and 6- carbon sugars rarely occur as straight chains. The carbon atoms with the aldehyde (or keto) group reversibly bonds to one of the other carbons by "sharing" a hydroxyl oxygen, forming a C-O-C linkage. This is known as an hemiacetal linkage. Typically, the result is a 5- or 6-member ring -- four or five carbon atoms plus the linking oxygen. A five-member form (e.g. a C1→C4 linkage) 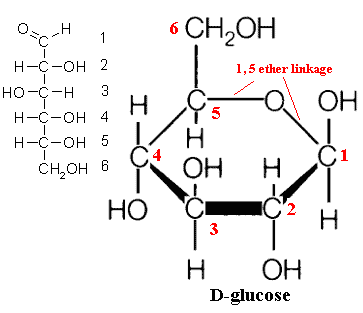 form is called a furanose. A six-member ring e.g. C1→C5 linkage) is a pyranose. A simple example, and perhaps the most common sugar monomer, is glucose. Its usual (pyranose) ring form is shown in the image. It can also occur as a furanose. In fact, the two forms are in equilibrium. Under biologically relevant conditions, the equilibrium so strongly favors the pyranose form of glucose that we can ignore the furanose. However, this is not necessarily the case for all sugars.
form is called a furanose. A six-member ring e.g. C1→C5 linkage) is a pyranose. A simple example, and perhaps the most common sugar monomer, is glucose. Its usual (pyranose) ring form is shown in the image. It can also occur as a furanose. In fact, the two forms are in equilibrium. Under biologically relevant conditions, the equilibrium so strongly favors the pyranose form of glucose that we can ignore the furanose. However, this is not necessarily the case for all sugars.
This is also the last time we will show the ring carbons. By the universal convention of biochemists, carbon atoms forming part of a ring structure are not shown with a 'C' symbol. They are simply indicated by the intersection of the bonds from the various groups (ligands) to which the carbon atom is attached. Very frequently, hydrogen ligands (H-) are not shown either. A line with nothing at the end means a hydrogen ligand, and an unlabelled intersection of bonds means a carbon atom. See examples below.
Sugar monomers are not always quite this simple. Each of the hydroxyl ligands is moderately chemically active, and all kinds of variants exist. An example, of particular relevance to fungi, is chitin. Chitin is a polymer of N-acetly-2-glucosamine, i.e., a glucose derivative in which the ligand CH3-CH2-NH- substitutes for the OH-group on C2. See the chitin glossary entry for an image.
In most of these examples, we have shown the structure of sugars using a Haworth Diagram. These are easy to draw and to understand, but they are rather crude tools because the bond angles are grossly distorted. Carbon normally forms tetrahedral structures, with the bonds about 108° apart. However, Haworth diagrams will do for our purposes, so long as we don't take them too seriously.
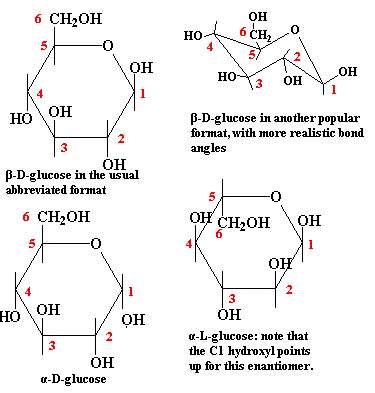 The figure above is labeled "D"-glucose for an important reason: it gives us an excuse to discuss three quick points about stereochemistry. Stereochemistry relates to the properties of compounds which are chemically identical, except that they are asymmetrical, and differ in the arrangement of ligands about one or more asymmetrical backbone atoms.
The figure above is labeled "D"-glucose for an important reason: it gives us an excuse to discuss three quick points about stereochemistry. Stereochemistry relates to the properties of compounds which are chemically identical, except that they are asymmetrical, and differ in the arrangement of ligands about one or more asymmetrical backbone atoms.
(1) Note that carbons 1 through 5 are asymmetrical in glucose. Each of these carbons is attached to four different ligands. Thus, the relative positions of the groups attached to the carbon atoms makes a difference. If, for example, we flipped the hydroxyl group on C2 so that it was above the ring, this would no longer be glucose. It would be mannose, a sugar with rather different chemical properties.
(2) If we took the mirror image of the entire molecule, all of the bonds would be in the same relative position. Thus we would have a molecule that ought to have exactly the same chemical properties as glucose, which it does -- sort of. The difficulty is that, when this reversed glucose interacts with some other asymmetrical biochemical, the two molecules no longer mesh in the same way. Consequently, we must distinguish between D-glucose and its mirror image (enantiomer), L-glucose. Don't worry about telling the difference. The biologically relevant form for sugars is usually the D-enantiomer. You can assume a figure shows the D-enantiomer unless someone tells you differently.
(3) C1 is a special case. In the linear form, C1 is not asymmetrical because it has only three ligands. However, when the C1 forms a pyranose linkage to C5, it becomes asymmetrical. In terms of our diagram, the -OH group on C1 might point down or up. Free glucose in solution is, once again, in equilibrium between the two forms, referred to as α- and β-D-glucose. These alternate forms of the hemiacetal are referred to as anomers. However, this time, neither form is strongly favored. (This is also not like the glucose-mannose example, since the two forms freely interconvert.) For free glucose, the exact form at any given time is unimportant. However, when glucose is linked to another sugar through the C1 hydroxyl group, the conformation becomes "frozen." Consequently, for glucose polymers, we need to distinguish between α (hydroxyl down) and β (hydroxyl up) linkages (glycoside bonds). Incidentally, the alpha-down/beta-up convention is reversed for L-enantiomers or, naturally enough, when the sugar monomer is represented upside-down.
General Features
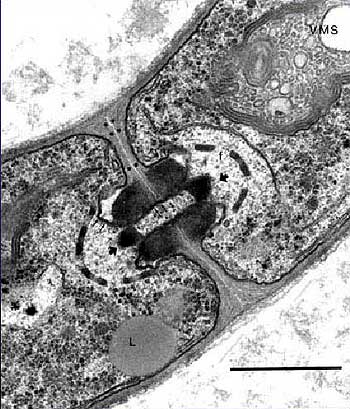 Fungal cells maintain a very high turgor pressure, so the integrity of the cell wall is a critical matter. Cabib et al. (2001). The composition of the fungal cell wall is rather variable. The variability appears to have phylogenetic significance, but few, to our knowledge, have followed that trail (but see Grun, 2003). In general, mycology has leapt directly from the ponderous fallacies of classical typological systematics to the facile, but sometimes equally fallacious, paradigms of molecular systematics. Consequently, there is remarkably little honest biology and biochemistry being applied to phylogenetic issues.
Fungal cells maintain a very high turgor pressure, so the integrity of the cell wall is a critical matter. Cabib et al. (2001). The composition of the fungal cell wall is rather variable. The variability appears to have phylogenetic significance, but few, to our knowledge, have followed that trail (but see Grun, 2003). In general, mycology has leapt directly from the ponderous fallacies of classical typological systematics to the facile, but sometimes equally fallacious, paradigms of molecular systematics. Consequently, there is remarkably little honest biology and biochemistry being applied to phylogenetic issues.
The situation is not improved by the usual non-specialist texts which characterize the fungal cell wall as a relatively simple structure made up of "cellulose" and chitin. Consider that the fungal cell wall can make up 30% or more of the dry weight of the fungus, and that the fungi are characterized by external digestion of food followed by selective absorption of the digestion products. Clearly, we can expect that the fungal cell wall will be a complex, specialized system.
It is all that; and, in addition, it is a highly dynamic system, constantly being regenerated and remodeled according to the needs of the moment. Adams 2004). Thus, many of the cell wall-associated proteins are enzymes whose function is to hydrolyze chitin and polysaccharides. The lesson is that this type of cell wall is, from a metabolic point of view, very different from insect exoskeletons or a plant cell walls, which are terminally differentiated structures.
Not unexpectedly, attempts to understand the biosynthesis of cell wall components have run into a maze of regulatory pathways which are difficult to sort out. García et al. (2004) applied brute force genomics methods to analyze gene responses to several different physical and chemical agents affecting cell wall integrity. The genetic responses in each case involved on the order of 100 different genes, with a significant different cohort of genes activated by each agent. Similarly, Lesage et al. (2004) identified 135 genes involved in the synthesis and regulation of the β-(1→3)-glucan component (see infra) alone (see also several similar studies cited by these authors). In fact, it has been estimated that 20% of the Saccharomyces genome is involved with cell wall biosynthesis. Durán & Nombela (2004). Some efforts are being made to pare these lists down to some "core" group of pathways. However, the magnitude of the problem has only become clear in the last few years, and it is much too early to say anything useful.
Structure
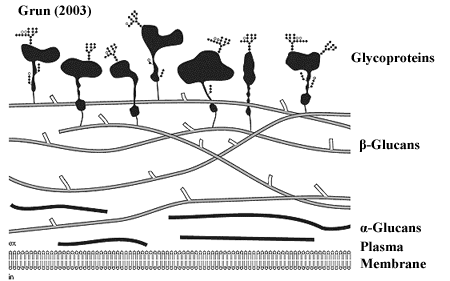 We include two diagrams of the fungal cell wall by Grün 2003) and Cabib et al. (2001). We've also thrown in Joan Miró's 1940) Chiffres et Constellations just because it has somewhat the same feel to it.
We include two diagrams of the fungal cell wall by Grün 2003) and Cabib et al. (2001). We've also thrown in Joan Miró's 1940) Chiffres et Constellations just because it has somewhat the same feel to it.
While each of these images speaks to us in its own way, we will work primarily with Grün's concept. The cell wall is generally constructed of three layers: (1) an α-glucan layer (a glucan is a polymer of glucose), (2) a β-glucan layer, and (3) an outer layer of glycoprotein. In addition, chitin may be a significant component of certain cell wall structures.
The α-glucan layer, if present, is generally composed of the α(1→3)-glucan polymer. However, α(1→4) glycosides are variably present. Compare glycogen, which is α(1→4)-glucan with (1→6) side chains. Where present, the α-glucan material appears as a fibrillar layer adjacent to the plasma membrane and is thought to serve a largely structural role, stiffening the basal layer of the cell wall.
The α-glucan layer is rarely represented in diagrams of the fungal cell wall because it does not occur in Saccharomyces, which is the usual model system. In fact, it has a rather peculiar phylogenetic distribution. Among ascomycetes, the alpha glucan is found in Schizosaccharomyces, but is not known from any other yeasts. The material is  common among all groups in the Pezizomycotina. However, in Lecanoromycetes, a very large proportion tends to be in the α(1→4) form. Alpha glucans also form a significant, sometimes even dominant, part of the cell wall in many basidiomycetes, but are completely absent outside the Hymenomycetes. Grün 2003). Although Schizosaccharomyces is often classified with the yeasts, its position is probably more basal. A number of studies show it branching with (paraphyletic) taphrinomycotines. See, e.g., Liu et al. (1999), An et al. (2002). We tend to prefer the methodology of these studies, which are neither biased by the superficial similarities of "yeast" forms nor confused by the usual problems with rDNA and mtDNA. Thus, it appears likely that the alpha glucan layer is primitive for all higher fungi, or at least for Ascomycota, with subsequent multiple losses.
common among all groups in the Pezizomycotina. However, in Lecanoromycetes, a very large proportion tends to be in the α(1→4) form. Alpha glucans also form a significant, sometimes even dominant, part of the cell wall in many basidiomycetes, but are completely absent outside the Hymenomycetes. Grün 2003). Although Schizosaccharomyces is often classified with the yeasts, its position is probably more basal. A number of studies show it branching with (paraphyletic) taphrinomycotines. See, e.g., Liu et al. (1999), An et al. (2002). We tend to prefer the methodology of these studies, which are neither biased by the superficial similarities of "yeast" forms nor confused by the usual problems with rDNA and mtDNA. Thus, it appears likely that the alpha glucan layer is primitive for all higher fungi, or at least for Ascomycota, with subsequent multiple losses.
The bulk material of the cell wall is usually in the form of β(1→3)-glucan. This forms a very stable hydrogen-bonded triple helix in solution, and probably in vivo. The packing of these triple helix structures appears to be controlled by the size and frequency of very short (1→6) side chains, sometimes consisting of only a single glucose monomer. Grün 2003). If so, this clearly provides a method for controlling the structure and conformation of the cell wall very simply and with very fine, localized control. However, essentially no work appears to have been done in this area. If anyone out there is looking for a potentially elegant and informative dissertation topic in a virtual research vacuum, this is it.
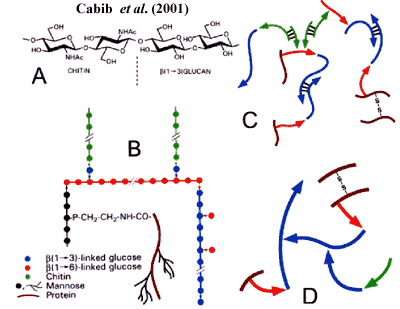 In addition to β(1→3)-glucan, the cell wall contains β(1→6)-glucan. We emphasize that this is not simply a β(1→3)-glucan with big side-chains, but a polysaccharide with a true β(1→6) backbone. This material may be peripheral to the bulk β(1→3)-glucan and is, in any case, strongly involved in cross-linking the various components of the cell wall, as shown in the figure from Cabib et al. (2001).
In addition to β(1→3)-glucan, the cell wall contains β(1→6)-glucan. We emphasize that this is not simply a β(1→3)-glucan with big side-chains, but a polysaccharide with a true β(1→6) backbone. This material may be peripheral to the bulk β(1→3)-glucan and is, in any case, strongly involved in cross-linking the various components of the cell wall, as shown in the figure from Cabib et al. (2001).
The outermost layer of the cell wall is composed of diverse proteins bearing polysaccharide side chains composed of mannose. The usual explanation is that these are attached through their mannan side chains via a (1→3) linkage with the β(1→6)-glucan. However, this is only a model. Real life appears to be very much more complex, involving a wide variety of different interactions between glycoproteins and bulk cell wall materials. Pitarch et al. (2002).
Finally, the fungal cell wall contains variable amounts of chitin. In many systems chitin is a major constituent of the cell wall. In others, it is involved only in cell division or reproductive structures and is virtually absent otherwise. Again. we are reluctant to say much about it, absent more detailed, phylogenetically-grounded studies of the actual ultrastructure in particular cases.
In general, the study of the fungal cell wall tends to be strong on models and somewhat weaker on data. One virtue of the brute force genomic and proteomic studies now being produced is that they clearly confront us with the scope of the problem. Fungal cells probably lack the diversity of metazoan tissues. However, each fungal cell must, for that very reason, be competent to perform a much wider variety of functions than a typical terminally-differentiated metazoan cell. Consequently, their superficial similarity and simplicity are likely to mask a very complex, plastic biochemical repertoire. Perhaps, after all, the Miró is the best representation, given the current state of our knowledge. ATW051113.
 form is called a furanose. A six-member ring e.g. C1→C5 linkage) is a pyranose. A simple example, and perhaps the most common sugar monomer, is glucose. Its usual (pyranose) ring form is shown in the image. It can also occur as a furanose. In fact, the two forms are in equilibrium. Under biologically relevant conditions, the equilibrium so strongly favors the pyranose form of glucose that we can ignore the furanose. However, this is not necessarily the case for all sugars.
form is called a furanose. A six-member ring e.g. C1→C5 linkage) is a pyranose. A simple example, and perhaps the most common sugar monomer, is glucose. Its usual (pyranose) ring form is shown in the image. It can also occur as a furanose. In fact, the two forms are in equilibrium. Under biologically relevant conditions, the equilibrium so strongly favors the pyranose form of glucose that we can ignore the furanose. However, this is not necessarily the case for all sugars. The figure above is labeled "
The figure above is labeled " Fungal cells maintain a very high
Fungal cells maintain a very high  We include two diagrams of the fungal cell wall by
We include two diagrams of the fungal cell wall by 
 In addition to β(1→3)-glucan, the cell wall contains β(1→6)-glucan. We emphasize that this is not simply a β(1→3)-glucan with big side-chains, but a polysaccharide with a true β(1→6) backbone. This material may be peripheral to the bulk β(1→3)-glucan and is, in any case, strongly involved in cross-linking the various components of the cell wall, as shown in the figure from
In addition to β(1→3)-glucan, the cell wall contains β(1→6)-glucan. We emphasize that this is not simply a β(1→3)-glucan with big side-chains, but a polysaccharide with a true β(1→6) backbone. This material may be peripheral to the bulk β(1→3)-glucan and is, in any case, strongly involved in cross-linking the various components of the cell wall, as shown in the figure from